Bioinformatics Analyses of Oct4 Pseudogenes
Anna Pang, Leah King, Zachary Jaloudi, Elisa Gubenko, Nneka Arinzeh, Anvi S. Bhatt and Lauren S. Sherman*
Rutgers New Jersey Medical School, Department of Medicine, Newark, NJ, USA
E-mail: shermala@njms.rutgers.edu
*Corresponding Author
Received 24 October 2025; Accepted 01 December 2025
Abstract
Oct4 is among the key genes involved in pluripotency. Oct4, through alternative splicing, produces at least three different transcripts, Oct4a, Oct4b, and Oct4b1. The Oct4a transcript has been focused on due to its role in pluripotency and multipotency of stem cells. There are several Oct4 pseudogenes with at least seven identified in the human genome. Some of these Oct4 pseudogenes have been found in various cancers. The Oct4 pseudogenes share high homology to the Oct4a transcript. There is limited information on the biologic role of Oct4 pseudogenes. More importantly, it is unclear how these pseudogenes affect the biology of the full-length transcript. Thus, it is important to understand these pseudogenes since their biology could lead to a better understanding of differentiation, cancer, and dedifferentiation to form cancer stem cells. This brief report used in silico analyses to analyze Oct4 and its pseudogenes. The implication for the findings on stem cells and cancer, as well as other related genes, are discussed.
Keywords: Stem cell, pseudogenes, Octamer 4, microRNA.
Introduction
Octamer-binding transcription factor 4 (Oct4), also known as Oct3 or POU5F1, is a transcription factor expressed in stem cells in adults, embryonic, and fetal tissues [1]. The role of Oct4 has been associated with pluripotency, cell proliferation, and self-renewal of embryonic stem and germ cells [1]. The literature indicates that Oct4 participates in the initiation and progression of a growing number of malignancies such as germ-cell tumors, bladder cancer, and liver cancer [1]. Furthermore, expression of Oct4 is required for maintaining the self-renewal and survival of cancer stem-like cells [1]. An area of research that requires significant attention involves non-coding RNAs and their role with the pseudogenes to regulate Oct4 in stem cells and cancers [1].
Pseudogenes comprise a class of non-coding RNAs, which are defined as genomic elements that resemble the genes [1]. Typically, pseudogenes are considered as non-functional genes or gene fragments due to their inability to translate functional or full-length proteins [2]. Pseudogenes are originally derived from functional genes, but exhibit some degenerative features such as inclusion of premature stop codons, deletions/inserts, or frameshift mutations [1, 2]. Pseudogenes that originate from reverse transcription of normal mRNA transcripts are labeled as processed pseudogenes and those that result from gene duplication are called non-processed pseudogenes [2]. The evidence indicate that pseudogenes might be important in regulating their originating genes [2]. Bioinformatics analyses have identified seven Oct4 pseudogenes [2]. Since some cancers also express high levels of Oct4, it is important for science to place greater consideration on Oct4 pseudogenes to determine how they might be important in controlling various cancers [3, 4].
Among the non-coding RNAs, micro RNAs (miRNAs) constitute an evolutionarily conserved class of pleiotropic small RNAs that function as a suppressor of gene expression post-transcriptionally [1]. miRNAs typically contribute to translational inhibition or mRNA degradation of large amounts of genes through their sequence-specific interactions with the 3’-untranslated regions (UTRs) of similar mRNA targets [1]. An miRNAs–Oct4 axis could be relevant to tumorigenesis [5]. This axis could be direct and indirect, by miRNA regulating other molecules to maintain cancer stem cells with high levels of Oct4 [6]. This short report provides informatic analyses to describe Oct4 pseudogenes and their potential roles in medicine with particular emphasis on cancer.
Materials and Methods
Sequence Coverage and Identity
The basic local alignment search tool (BLAST) is a popular sequence analysis tool that could identify short matches between two sequences with alignments from hot spots [7]. Several types of BLAST programs exist to compare all combinations of nucleotide or protein queries with nucleotide or protein databases [7]. Beyond the capability to performing alignments, BLAST provides statistical information that contributes to deciphering the biological significance of the alignment using an expect value or a false-positive rate [7]. Known Oct4 transcripts and pseudogenes (Supplemental Table 1) were aligned using default parameters on the National Center for Biotechnology Information (NCBI) platform to identify their query coverage and sequence identity relative to the parent Oct4a.
Sequence Alignment
A multiple sequence alignment program for proteins produces biologically meaningful multiple sequence alignments of divergent sequences [8–10]. Cladograms or phylograms are available to view the evolutionary relationships of a protein [8–10]. Known Oct4 transcripts and pseudogenes have been aligned using default parameters on the European Bioinformatics Institute (EBI) platform. Clustal Omega was used for local sequence alignment, and Needle was used for global sequence alignment.
Exon Identification
Ensembl is a system that generates genomic datasets through a system that is designed to analyze, store and distribute data, and interpret through open data release from 87 species [11]. The database was queried for Oct4 transcripts and pseudogenes to identify each sequence’s chromosomal loci and exon positions.
Predicted miRNA Binding Identification
miRBase is a microRNA database used to predict microRNA-transcript binding. Each Oct4 transcript and pseudogene was queried for predicted miRNA binding sites. Top miRNA hits for each sequence were compared to determine whether the miRNA sequences may bind with other Oct4 sequences.
MOTIF Conservation
MOTIF Search (https://www.genome.jp/tools/motif/) allows querying of multiple sequences for conserved motifs. The Oct4 transcripts and pseudogenes were queried using default parameters.
Results
Features of Oct4 Pseudogenes Relative to Oct4 Transcripts
Pseudogenes have been shown to regulate related full-length genes [2]. NCBI BLAST compared different Oct4 pseudogenes with the full length Oct4 transcripts (Tables 1 and 2). The identified seven Oct4 pseudogenes showed high sequence homologies (>90%) with the Oct4 gene (Tables 1 and 2). We conducted multiple sequence alignment and global sequence alignments of the Oct4 pseudogenes and the Oct4 gene using Clustal Omega and Needle, respectively (Supplemental Tables S2 and S3a-g).
The Oct4 gene can generate at least three transcripts: Oct4a, Oct4b, and Oct4b1 [12]. The subsequent analyses focus on the Oct4a transcript due to its role in pluripotency of healthy and cancer stem cells [12, 13]. Furthermore, Oct4b does not seem to have a role in sustaining stem cell self-renewal, although it might be able to respond to cell stresses [12].
Table 1 Oct4 pseudogenes relative to Oct4a
| Pseudogene | Chromosome | Query cover to Oct4a (%) | Sequence identity to Oct4A (%) |
| Oct4-pg1 | 8 | 85% | 98% |
| Oct4-pg2 | 8 | 93% | 80% |
| Oct4-pg3 | 12 | 100% | 98% |
| Oct4-pg4 | 1 | 100% | 97% |
| Oct4-pg5 | 10 | 100% | 88% |
| Oct4-pg6 | 3 | 27% | 83% |
| Oct4-pg7 | 3 | No significant similarity found | |
| The data were obtained from NCBI using BLAST [14]. | |||
Table 2 Pseudogenes relative to Oct4B and Oct4B1
| Query cover | Sequence identity | Query cover | Sequence identity | |
| Pseudogene | to Oct4B (%) | to Oct4B (%) | to Oct4B1 (%) | to Oct4B1 (%) |
| Oct4-pg1 | 57% | 98% | 7% | 99% |
| Oct4-pg2 | 80% | 81% | 10% | 87% |
| Oct4-pg3 | 62% | 99% | 11% | 98% |
| Oct4-pg4 | 62% | 98% | 11% | 98% |
| Oct4-pg5 | 100% | 88% | 13% | 95% |
| Oct4-pg6 | 27% | 83% | 4% | 93% |
| Oct4-pg7 | No significant similarity found | No significant similarity found | ||
| The data were obtained from NCBI using BLAST [14]. | ||||
Exons of Oct4 Pseudogenes and Oct4 Transcripts
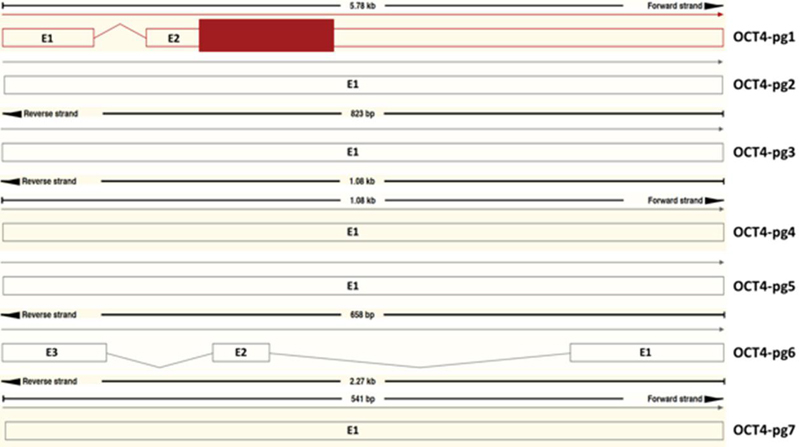
The nucleotide sequences of Oct4-pg1, Oct4-pg3, and Oct4-pg4 were shown to be similar to Oct4a transcript (POU5F1-001) (Table 1). Comparison of Oct4a with the Oct4 pseudogenes identified five Oct4a exons within Oct4-pg1, Oct4-pg3, and Oct4-pg4 [15]. Oct4-pg5 transcript lacks Exon 1 and Oct4-pg7 lacks Exons 1 and Exon 4 as well as part of Exon 2; Oct4-pg2 has part of Exon 5, and OCT4-pg6 has the five exons incompletely [15]. Upon further investigation using Ensembl, the information regarding these exons seem to show inconsistency (Tables 3 and 4, Figures 1 and 2). Interestingly, Oct4-pg1, Oct4-pg3, and Oct4-pg4 may have the ability to generate proteins [15]. In total, the analyses mostly show weak evidence for protein expression by Oct4 pseudogenes.
Table 3 Exons of Oct4 pseudogenes
| Exon(s); | |||
| Pseudogene | Chromosome | coding exon(s) | Exon Ensembl ID |
| Oct4-pg1 | 8 | 2; 1 | ENSE00001852952 |
| ENSE00001814817 | |||
| Oct4-pg2 | 8 | 1; 0 | ENSE00002116992 |
| Oct4-pg3 | 12 | 1; 0 | ENSE00001717808 |
| Oct4-pg4 | 1 | 1; 0 | ENSE00001646566 |
| Oct4-pg5 | 10 | 1; 0 | ENSE00001694027 |
| Oct4-pg6 | 3 | 3; 0 | ENSE00001892786 |
| ENSE00001934799 | |||
| ENSE00001930279 | |||
| Oct4-pg7 | 3 | 1; 0 | ENSE00002087477 |
| These data were obtained from Ensembl [11]. | |||
Table 4 Exons of POU5F1 (Oct4) transcripts
| Name | Chromosome | Exon(s); coding exon(s) | Exon Ensembl ID |
| POU5F1-001 | 6 | 5; 5 | ENSE00001834753 |
| ENSE00003605759 | |||
| ENSE00003631186 | |||
| ENSE00003697734 | |||
| ENSE00003736761 | |||
| POU5F1-002 | 6 | 4; 3 | ENSE00002568331 |
| ENSE00003606772 | |||
| ENSE00003697734 | |||
| ENSE00002055331 | |||
| POU5F1-003 | 6 | 2; 0 | ENSE00001891266 |
| ENSE00001843145 | |||
| POU5F1-004 | 6 | 5; 3 | ENSE00002243764 |
| ENSE00003566428 | |||
| ENSE00003606772 | |||
| ENSE00003697734 | |||
| ENSE00003736761 | |||
| POU5F1-005 | 6 | 3; 3 | ENSE00002033137 |
| ENSE00003697734 | |||
| ENSE00003702101 | |||
| POU5F1-006 | 6 | 5;3 | ENSE00002043181 |
| ENSE00003566428 | |||
| ENSE00003606772 | |||
| ENSE00003697734 | |||
| ENSE00003702101 | |||
| POU5F1-007 | 6 | 5; 4 | ENSE00002043181 |
| ENSE00003702358 | |||
| ENSE00003631186 | |||
| ENSE00003697734 | |||
| ENSE00003745997 | |||
| POU5F1-201 | 6 | 6; 4 | ENSE00003750435 |
| ENSE00003730318 | |||
| ENSE00003742006 | |||
| ENSE00003713303 | |||
| ENSE00003723189 | |||
| ENSE00003744866 | |||
| POU5F1-202 | 6 | 5; 4 | ENSE00003751524 |
| ENSE00003702358 | |||
| ENSE00003631186 | |||
| ENSE00003697734 | |||
| ENSE00003727125 | |||
| The information was obtained from Ensembl [11]. | |||
Figure 1 Exons of Oct4 pseudogenes. The diagram was developed with Ensembl [11].
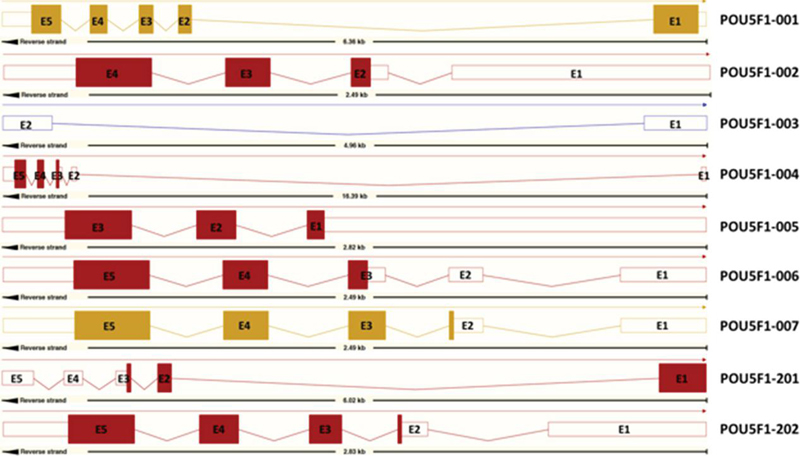
Figure 2 Exons of Oct4 transcripts. These data were obtained from Ensembl [11].
Potential Interacting Sites of miRNA and Consensus Sequences
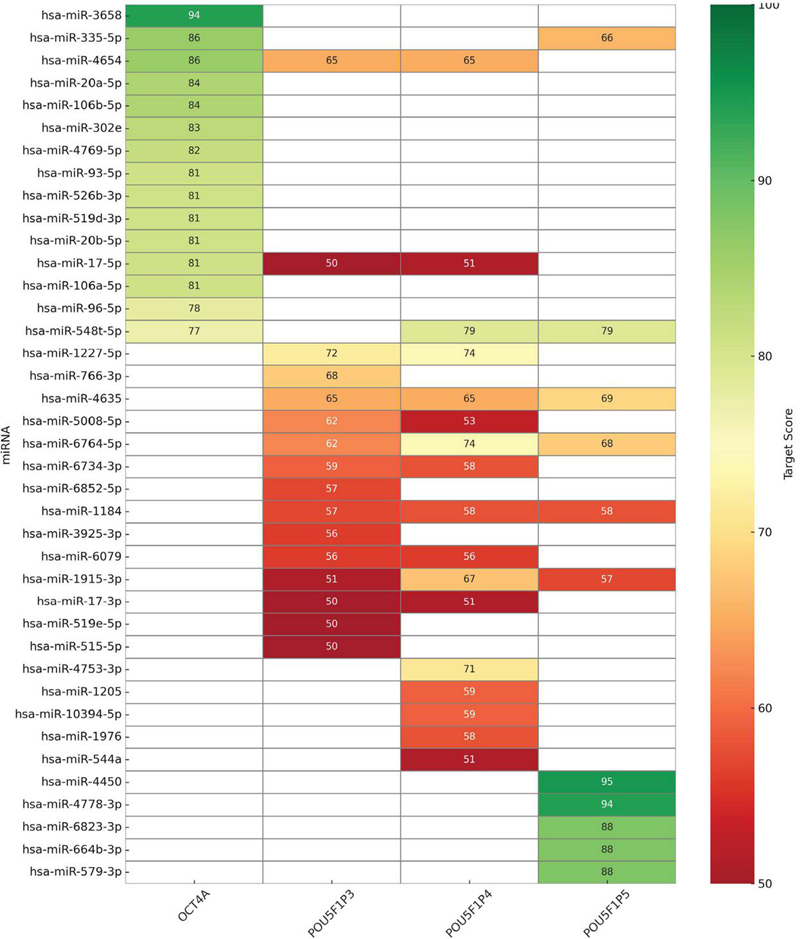
We employed a more stringent alignment with Oct4a that is linked to healthy and cancer stem cells. We observed a significant alignment with pseudogenes 3-5 (Table S2). Since miRNA can suppress translation, we asked if these pseudogenes could sponge miRNA to prevent them from affecting Oct4a translation. To this end, we analyzed the top miRNAs that could interact with Oct4a and the pseudogenes (Figure 3). Although there were potential miRNA interacting sites on the transcripts, there was no clear indication of the degree of competition of the miRNA.
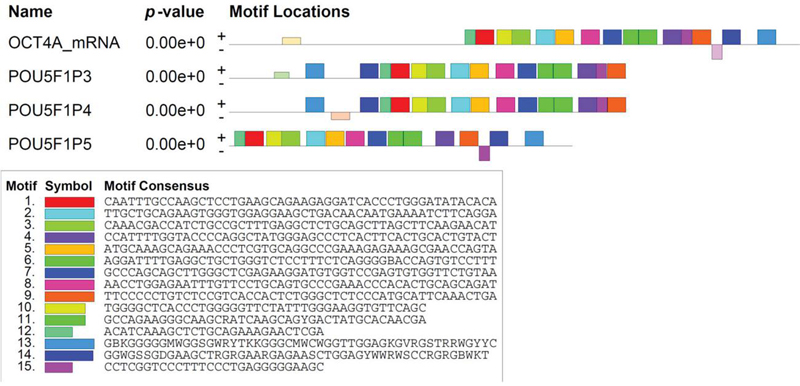
Although this analysis did not examine the pseudogenes as potential sponges for transcription factors, we examine them, along with Oct4a for consensus sequences. The results showed several share sequences (Figure 4). The relevance of the information is discussed below.
Figure 3 Top aligned miRNA with Oct4A and pseudogenes.
Figure 4 Consensus sequences in Oct4a and pseudogenes.
Discussion
The Oct4 transcription factor plays a key role in maintaining stem cell pluripotency [16]. The evidence indicated that, in addition to embryonic and adult healthy stem cells, Oct4 is also relevant to cancer stem cell [13, 15]. The information on Oct4 would be improved with careful studies to discern the full-length transcript with pseudogenes. This study will allow for proper selection of primers to identify the various pseudogenes from the full-length transcript. RT-PCR artifacts and misdetection of the Oct4A isoform could be partly derived by the amplification of highly homologous Oct4 pseudogenes at the transcript level. There is a possibility that Oct4 pseudogenes may have some functional activity at the transcript or protein level in tumor cells and tissues [15]. In tumors, it is interesting that Oct4 pseudogenes expression seem to be differentially expressed, depending on the tumor [15]. The full-length and pseudogenes of Oct4 could be important to the biology of tumors.
The report identified a new layer of post-transcriptional regulation of Oct4 [1]. The study reported on abnormal activation of Oct4-pg4 in hepatocellular carcinoma. The expression level of Oct4-pg4 positively correlated with Oct4. Furthermore, the authors found that Oct4 transcript can be directly targeted by tumor-suppressive micro RNA-145 (miR-145) [1]. Downregulation of Oct4-pg4 leads to an increased availability of miR-145 to bind Oct4 transcript, resulting in decreased Oct4 protein [1]. Overexpression of Oct4-pg4 leads to decrease in free miR-145, leading to increase of Oct4 protein [1]. Based on this, the authors premise that Oct4-pg4 functions as a natural miRNA sponge to protect Oct4 translation [1]. Thus Oct4-pg4 is implicated as having an oncogenic role in hepatocarcinogenesis as it can promote growth and tumorigenicity of hepatocellular carcinoma [1]. Researchers have suggested a conserved miR-145 binding site for almost all of the Oct4 pseudogenes [15]. The wide expression of Oct4 pseudogenes in different types of cancer may be associated with tumorigenesis when considering the role of miR-145 in tumor suppression [15].
There are several shared consensus sequences of Oct4a and the pseudogenes (Figure 4). Although not addressed in this study, it would be interesting to determine if those sequences can bind to miRNAs. Figure 3 shows potential miRNA that could be used in functional studies to understand if indeed the pseudogenes are indeed miRNA sponges, similar to circular RNA [17].
The Oct4 gene, although linked to stem cells, is involved in cancer pathogenesis, including drug resistance and cancer stem cells [15, 18, 19]. Thus, it is important to further discuss how this report could impact studies on tumorigenesis. Oct4 pseudogenes are differentially expressed in various types of tumors cell types as well as in human pluripotent stem cells [15].
The evidence is unclear with respect to the ability of the pseudogenes to be translated to protein. This study indicated that Oct4-pg1 might produce protein with little evidence of translation by the other pseudogenes. The unanswered question regarding translation of the pseudogenes indicates a more in-depth analysis of this function, as well as a potential sponging effect, as reported for miR-145. This study also requires a function studies to determine how the informatics analyses regulate Oct4 as well as other stem cell genes such as Nanog [20].
References
[1] Wang, L., Z.-Y. Guo, R. Zhang, B. Xin, R. Chen, J. Zhao, T. Wang, W.-H. Wen, L.-T. Jia, and L.-B. Yao. 2013. Pseudogene OCT4-pg4 functions as a natural micro RNA sponge to regulate OCT4 expression by competing for miR-145 in hepatocellular carcinoma. Carcinogenesis 34: 1773–1781.
[2] Suo, G., J. Han, X. Wang, J. Zhang, Y. Zhao, Y. Zhao, and J. Dai. 2005. Oct4 pseudogenes are transcribed in cancers. Biochem Biophysical Res Commun 337: 1047–1051.
[3] Kolenda, T., P. Chałaj, A. Cichowicz, A. Trojańska, A. Bałoniak, M. Kwaśniewska, M. Odrobińska, K. Guglas, J. Kozłowska-Masłoń, P. Gieremek, P. Poter, M. Janiczek-Polewska, A. Florczak-Substyk, A. Przybyła, P. Mantaj, K. Regulska, B. J. Stanisz, Z. Cybulski, and U. Kazimierczak. 2025. Pseudogenes in the carcinogenesis: epithelial-to-mesenchymal transition process and cancer initiating cells. Rep Pract Oncol Radiother 30: 366–384.
[4] Sharma, D., S. Gupta, G. Koshy, V. K. Sharma, M. Kamboj, and A. Hooda. 2025. Expression profile of cancer stem cell markers SOX2, OCT4 & NANOG in salivary gland malignancies: A systematic review. Indian J Med Res 161: 636–646.
[5] Bliss, S. A., G. Sinha, O. A. Sandiford, L. M. Williams, D. J. Engelberth, K. Guiro, L. L. Isenalumhe, S. J. Greco, S. Ayer, M. Bryan, R. Kumar, N. M. Ponzio, and P. Rameshwar. 2016. Mesenchymal Stem Cell-Derived Exosomes Stimulate Cycling Quiescence and Early Breast Cancer Dormancy in Bone Marrow. Cancer Res 76: 5832–5844.
[6] Sinha, G., A. I. Ferrer, S. Ayer, M. H. El-Far, S. H. Pamarthi, Y. Naaldijk, P. Barak, O. A. Sandiford, B. M. Bibber, G. Yehia, S. J. Greco, J. G. Jiang, M. Bryan, R. Kumar, N. M. Ponzio, J. P. Etchegaray, and P. Rameshwar. 2021. Specific N-cadherin-dependent pathways drive human breast cancer dormancy in bone marrow. Life Sci Alliance 4.
[7] McGinnis, S., and T. L. Madden. 2004. BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res 32: W20–W25.
[8] Li, W., A. Cowley, M. Uludag, T. Gur, H. McWilliam, S. Squizzato, Y. M. Park, N. Buso, and R. Lopez. 2015. The EMBL-EBI bioinformatics web and programmatic tools framework. Nucleic Acids Res 43: W580–W584.
[9] McWilliam, H., W. Li, M. Uludag, S. Squizzato, Y. M. Park, N. Buso, A. P. Cowley, and R. Lopez. 2013. Analysis tool web services from the EMBL-EBI. Nucleic Acids Res 41: W597–W600.
[10] Sievers, F., A. Wilm, D. Dineen, T. J. Gibson, K. Karplus, W. Li, R. Lopez, H. McWilliam, M. Remmert, and J. Söding. 2011. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Systems Biol 7: 539.
[11] Yates, A., W. Akanni, M. R. Amode, D. Barrell, K. Billis, D. Carvalho-Silva, C. Cummins, P. Clapham, S. Fitzgerald, L. Gil, C. G. Girón, L. Gordon, T. Hourlier, S. E. Hunt, S. H. Janacek, N. Johnson, T. Juettemann, S. Keenan, I. Lavidas, F. J. Martin, T. Maurel, W. McLaren, D. N. Murphy, R. Nag, M. Nuhn, A. Parker, M. Patricio, M. Pignatelli, M. Rahtz, H. S. Riat, D. Sheppard, K. Taylor, A. Thormann, A. Vullo, S. P. Wilder, A. Zadissa, E. Birney, J. Harrow, M. Muffato, E. Perry, M. Ruffier, G. Spudich, S. J. Trevanion, F. Cunningham, B. L. Aken, D. R. Zerbino, and P. Flicek. 2016. Ensembl 2016. Nucleic Acids Res 44: D710–716.
[12] Wang, X., and J. Dai. 2010. Concise review: isoforms of OCT4 contribute to the confusing diversity in stem cell biology. Stem Cells 28: 885–893.
[13] Patel, S. A., S. H. Ramkissoon, M. Bryan, L. F. Pliner, G. Dontu, P. S. Patel, S. Amiri, S. R. Pine, and P. Rameshwar. 2012. Delineation of breast cancer cell hierarchy identifies the subset responsible for dormancy. Sci Rep 2: 906.
[14] Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J Mol Biol 215: 403–410.
[15] Poursani, E. M., B. M. Soltani, and S. J. Mowla. 2016. Differential expression of OCT4 pseudogenes in pluripotent and tumor cell lines. Cell J (Yakhteh) 18: 28.
[16] Greco, S. J., K. Liu, and P. Rameshwar. 2007. Functional similarities among genes regulated by OCT4 in human mesenchymal and embryonic stem cells. Stem Cells 25: 3143–3154.
[17] Kulcheski, F. R., A. P. Christoff, and R. Margis. 2016. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J Biotechnol 238: 42–51.
[18] Villodre, E. S., F. C. Kipper, M. B. Pereira, and G. Lenz. 2016. Roles of OCT4 in tumorigenesis, cancer therapy resistance and prognosis. Cancer treatment reviews 51: 1–9.
[19] Zhang, Q., Z. Han, Y. Zhu, J. Chen, and W. Li. 2020. The role and specific mechanism of OCT4 in cancer stem cells: a review. Intl J Stem Cells 13: 312–325.
[20] Booth, H. A. F., and P. W. Holland. 2004. Eleven daughters of NANOG. Genomics 84: 229–238.
International Journal of Translational Science, Vol. 2, 165–176
doi: 10.13052/ijts2246-8765.2025.024
© 2026 River Publishers