Detection of Rice Plant Disease Using Deep Learning Techniques
S. Babu1, M. Maravarman2,* and R. Pitchai2
1Department of Computing Technologies, SRM Institute of Science and Technology, Kattankulathur, Chennai, India
2Department of Computer Science and Engineering, B V Raju Institute of Technology, Narsapur, Medak, Telangana, India
E-mail: babubalaji2k5@gmail.com; maravarman.it@gmail.com; pitchrks1984@gmail.com
*Corresponding Author
Received 30 September 2021; Accepted 16 November 2021; Publication 22 January 2022
Abstract
Deep learning has recently grown a lot of interest as a way to create a fast, efficient, and reliable image identification and categorization system. India, being one of the world’s most important rice producers and consumers, relies heavily on rice to propel its economy and provide its food needs. In the crop protective device, early and precise diagnosis of plant diseases is critical. Traditionally, identification was done either through visual inspection or laboratory testing. It is critical to identify any disease early and perform the necessary treatment to the damaged plants in order to guarantee the rice plants’ healthy and proper growth. Because disease detection by hand takes a long time and requires a lot of effort, having an automated system is unavoidable. A rice plant disease identification method depends on deep learning methodologies are presented in this research. Leaf smut, bacterial leaf blight, sheat blight, and brown spot diseases are four of the most frequent rice plant diseases identified in this study. The rice plant disease is identified and recognized using deep learning algorithms. This method of early detection of rice diseases could be utilized as a preventative tool as well as an early detection. The proposed approach provides enhanced accuracy of 99.45% and it is compared with the existing state-of-the-art approaches.
Keywords: Rice plant disease, classification, pattern recognition, convolutional neural network (CNN).
1 Introduction
Agriculture is a major source of revenue in many countries across the world. Based on the importance of agriculture, farmers choose their crops, paddies, and pesticides to help the plant develop faster in a limited amount of time. Plant diseases play a role in both covert and overt environmental impact. As these diseases proliferate over the world, they harm the plant’s overall function as well as the financial situation by drastically reducing the amount of crops grown. Rice is consumed by half of the earth’s population on a regular basis. According to the World Economic forum, rice consumption is expected to grow by 51% by 2025. Numerous factors, particularly mineral deficiency, disease prediction have an impact on rice productivity. Early identification and categorization of crop plant diseases is one among the most important actions in agricultural practices. Every year, virus infection causes a significant financial damage to farmers [1]. As a result, early, accurate, and prompt detection of the condition avoids product loss while also improving product quality. Smart farming strategies are used to manage productivity, food and nutrition security, efficiency, and environmental effect. Food production plays an increasingly important role as the world’s population grows. The quality of food products is critical for maintaining optimum health in humans. It is necessary to conduct an examination of many factors that contribute to agricultural productivity loss in order to provide this healthy environment. Identification of the afflicted portion, Image classification, detection of an abnormality in plant leaves, and image analysis in agriculture are examples of open large area research. As the world’s population grows, so do the issues that agriculture faces on a daily basis [2, 3]. In agriculture, land speculation and water scarcity are critical.
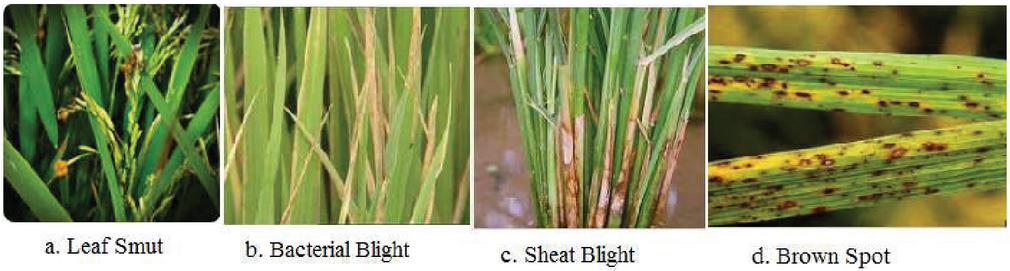
Figure 1 Four diseases of rice plant.
Modern agriculture employs a variety of technology to address these issues in real time. Deep learning is defined as a burgeoning branch of machine learning. The deep learning research methodology has been widely and productively utilized in a variety of fields. It’s a group of machine learning approaches that use several stages of non-linear processing of information for unsupervised and supervised extraction of features and modification, as well as pattern recognition and categorization. Deep learning is a technique in which algorithms are organized in layers to construct a convolution neural networks that can understand and make choices by itself. Every nation needs to concentrate on its manufacturing breakthroughs to keep up with the 4th industrial revolution that will include embedded technologies that can make choices without personal communication. The authors of this research concentrated on the detection of four rice leaf disease identification that are Leaf smut, bacterial leaf blight, sheat blight, and brown spot [4, 5]. The occurrence of these diseases in most nations is the basis for choosing these disorders. If not addressed as soon as possible, this could be disastrous for production quantity and quality, as well as overall production. Machine Learning principles, instead of visual observations and identification, can surely aid with this goal’s effectiveness. The characteristics of these diseases are specified as below. The Figure 1 denotes the various four diseases of rice plant.
• Leaf Smut
False Smut of rice is caused by this disease, which decreases grain yield and quality. The disease is found in over 40 nations, primarily in Asia’s rice-producing countries.
• Bacterial Blight
Rice bacterial blight, or simply rice blight, is indeed a deadly bacterium that causes that one of the most destructive diseases to rice varieties. In extreme outbreaks, yield reduction can exceed 75%, and a million acres of land of rice are affected every year.
• Sheat Blight
Rhizoctonia solani causes sheath blight, a fungal disease. Early tillers could also be killed if infected plants senesce or dried out and decay more quickly. As a result, the disease has the potential to drastically lower the canopy’s number of leaves.
• Brown Spot
Brown spot disease in rice is caused by the fungus Cochliobolus miyabeanus. The sickness that caused the Bengal famine in 1943 was caused by this disease. During World War II, the United States explored using it as a bioweapon against Japan.
Monitoring disease incidences and frequencies is critical for early discovery of afflicted plants, prompt treatment, and, most significantly, the development of future disease prevention techniques to reduce losses [6]. Crop disease monitoring in Bangladesh has traditionally been done by manually detecting any abnormality in plants, having experts classify that abnormality as a disease, and then proposing proper treatment. The major contributions of the proposed work are
• An approach to design a efficient rice plant disease prediction
• Three various rice disease classification has to be done effectively
• To enhance the novelty of the proposed approach deep learning methodology is included for better accuracy in disease prediction.
• The efficiency of the proposed approach is compared with the state-of-the-art approaches.
2 Related Works
Mahanty et al. [7] proposed using deep learning to create a smartphone based disease detection approach. Using collections of 54,200 photos of good and infected plant leaves, they employed CNN to construct the required model. Using photos, CNN was taught to recognise 14 crop varieties and 20 infected species They assessed CNN’s suitability for the categorization of plants, crops, and illnesses. AlexNet and GoogLeNet were the two architectures they used. Their model had a 99.35 percent accuracy rate. Despite the fact that their model produced state-of-the-art results, it performed badly when evaluated on a set of photos shot under various conditions. Barbedo et al. [8] conduct research on the major aspects that influence the design and efficiency of deep neural network models used in plant pathology. An in-depth examination of the subject, highlighting its benefits and drawbacks, should bring to more reasonable judgments on the issue. Rathore et al. [16] proposed a deep learning CNN model to automatically detect and predict the rice plant diseases. Since manual detection and prediction of disease is much time consuming automatic detection based on computer aided techniques are needed. For the detection of rice nitrogen deficit.
Sethy et al. [17] presents a CNN based technique. By substituting the very last output layer of CNN with such a pre-eminent predictor like SVM, the pre-trained Framework is adjusted to increase classification accuracy. With 5790 imaging examples, six top deep learning architectures are utilised to detect nitrogen defect: ResNet-18, ResNet-50, GoogleNet, AlexNet, VGG-16, and VGG-19 with SVM. The accuracy, specificity, precision, false positive rate (FPR), and F1 measure of every predictor are all tested and compared. Rangarajan et al. [9] provide a method for recognising weeds and species of plants using coloured graphics in their study. In their research, they used CNN, which was evaluated on a total of 10,413 photos containing 22 plants and crop varieties. The CNN model proved able to reach an accuracy of 86.2 percent in categorization. The network struggled to classify several plant species, which is thought to be due to a lack of training images for those living things. The authors wanted to use Deep Learning techniques for detecting plant diseases, that would allow consumers to identify diseases rapidly and effortlessly, allowing them to take appropriate action at an early stage. They tested 2589 original photos and trained their model with 30880 images using the Caffe deep learning approach [10]. Again, using the outcomes of 100 individual simulations, data analysis is used to select the best categorization model. ResNet-50SVM outperformed the other five CNN-based classification techniques in descriptive statistics, with an accuracy of 99.84 percent.
For quicker and improved search facility, a new adaptive approach that integrates genetic algorithms (GA) and particle swarm optimization (PSO) is developed in Moslehi et al. [18]. The hybrid algorithm combines the benefits of both the PSO and GA approaches. The primary purpose of the feature selection procedure is to choose a subset of exceptional and crucial characteristics by removing irrelevant features and the set’s lower amount of data. This decrease in dimensions aids in the speed of the learning process, as well as the creation of simple and understandable forecasting models and the avoidance of over fitting. Matin et al. [18] used the AlexNet technique for detecting three common rice plant disease: bacteria blight, leaf spots, and leaf smut, and the results were far superior to earlier studies. AlexNet is a supervised neural classification system with a twist. Due to the use of an effective approach and picture augmentation, this work achieves a level of accuracy of above 99 percent. Patidar et al. [20] proposed a paradigm to define the rice plant disease prediction using deep residual learning. Using images of contaminated rice plants, the subsequent study proposes a paradigm for detecting and classifying infections in rice plants, one of its most important crops in the Indian staple diet. Bacterial leaf blight, Brown spot, and Leaf smut were the three disorders that were mostly studied. The UCI Machine Learning Repository’s Rice Plant Leaf Collection has been used.
3 Proposed Methodology
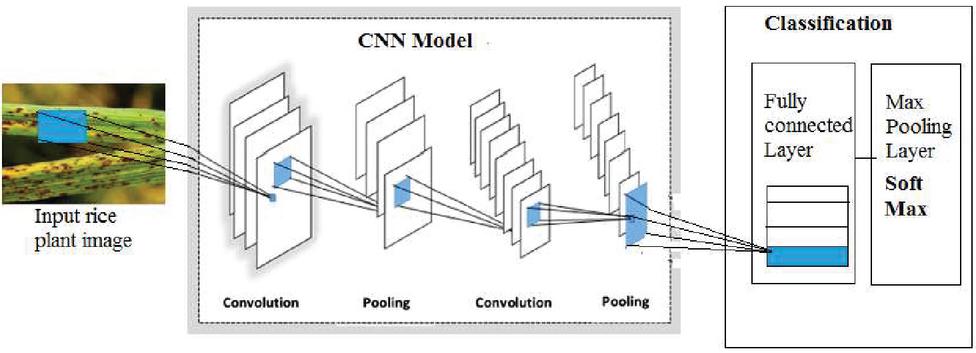
To date, many modeling approaches such as neural networks and multiple linear regressions have been used to forecast illness in plant communities. However, new forecasting software is needed to better grasp plant-pathogen-environment connections due to their incapacity to anticipate the value of unexpected data sets and lengthier training timeframes. Following it, a variety of architectures were created. A state-of-the-art convolution neural network is evaluated and fine-tuned for the job of plant disease recognition and characterization using photos from PlantVillage in this work. PlantVillage has 54,306 photos for 14 agricultural plants, with 26 illnesses. Colored photos of various sizes were used to create the graphics. For VGG net, ResNet, and DenseNets designs, the photos are first downsized to 224224 pixels. Images for the Inception V4 architecture, on either hand, are downsized to 299 299 pixels. To ensure all pixel values consistent with the network’s initial values, data is computed by dividing them by 255. The dataset is divided into 20%. The proposed methodology is shown in the Figure 2.
Figure 2 Proposed Deep Learning Model for disease identification.
3.1 Resnet
ResNet is a network-in-network (NIN) design that uses layered residual units to create a network within a network. The set of foundation blocks utilised to form the network is made up of these leftover units. The ResNet Architecture is composed of a collection of residual units that serve as construction pieces. Convolution, pooling, and layering make up the residual units. The architecture resembles that of the VGG network. The use of global average pooling instead of fully-connected layers is to blame for this. ResNet was updated again to improve accuracy by changing the residual module to employ identification assignments.
3.2 Inception Model
A pooling layer and convolution layers are placed together in the Inception module. The convolutions range in size from 11 to 33% to 55%. The usage of a bottleneck layer, that is 11 convolutions, is another notable element of the Inception module. The bottleneck layer aids in lowering computation demands [11]. Inside the module, there is also a pooling layer that is utilized for feature reduction. Inception v4 uses residual connections to substitute the filter concatenation phase of the proposed architecture. The GoogLeNet Inception V4 was fine-tuned utilizing pre-trained values from ImageNet. On the upper layer, reduction and construction of a current design with Average pooling layer (88) was conducted, as well as dropping and softmax.
3.3 VGC Net
For the ILSVRC-2014 competition, VGG net was created as a CNN model. On the validation set, the model had a top-five error rate of 7.5 percent, which earned them second position in the challenge. With only 33 convolutional layers put on top of one another in increasing thickness, the model is usually typified by its simplicity.
3.4 DenseNet
Densenet is a fully convolutional structure with a lot of connections. All layers in the system are connected directly to one another in a feed-forward manner that enables optimal flow of information between them. All previous layers’ functionality are utilised as inputs through every layer, and its own function are used as sources into all subsequent layers. DenseNets solves the back to a simpler while drastically reducing the number of variables.
4 Results and Discussions
This research demonstrates the necessity of rice plant disease detection in today’s world. This model was created in Python using Deep Learning. Pictures from the PlantVillage dataset were utilized for 80% of the training and 20% for testing the model’s accuracy. There are 38 different classes represented in these photos. For testing, 20% of each class was chosen at random. There were also some real-time photographs used. The accuracy measure and categorized loss are being used to evaluate the models in each experiment. Every model’s effectiveness is visually shown with accuracy and loss [12, 13]. The outcome of the given method is determined by computing an actual loss score and accuracy based on test dataset. The suitability of a state-of-the-art deep learning model for the assignment of infected plant diagnosis using photographs was assessed in this approach. The results of the proposed approach are specified in the Table 1.
Table 1 Results of the proposed approach
| Layers | Training | Training | Validation | Validation | |
| Model | Used | Accuracy (%) | Loss (%) | Accuracy (%) | Loss (%) |
| Resnet | 101 | 99.92 | 0.0230 | 99.84 | 0.0211 |
| Inception Model | 16 | 99.83 | 0.0197 | 99.62 | 0.0191 |
| VGC net | 16 | 90.04 | 0.0531 | 90.81 | 0.0623 |
| Densenet | 121 | 99.74 | 0.0121 | 99.17 | 0.0103 |
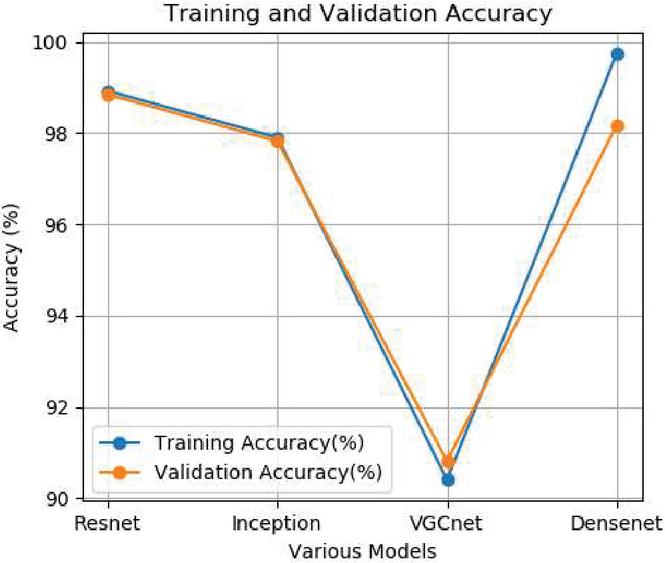
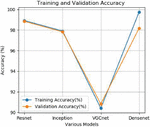
Figure 3 Training and validation accuracy.
In this study after experimenting different runs the training and validation accuracy as well as training and testing loss are analyzed. The average training and validation accuracy for Resnet is 99.88%, Inception model is 99.72%, VGCnet is 90.42% and Densenet is 99.45%.
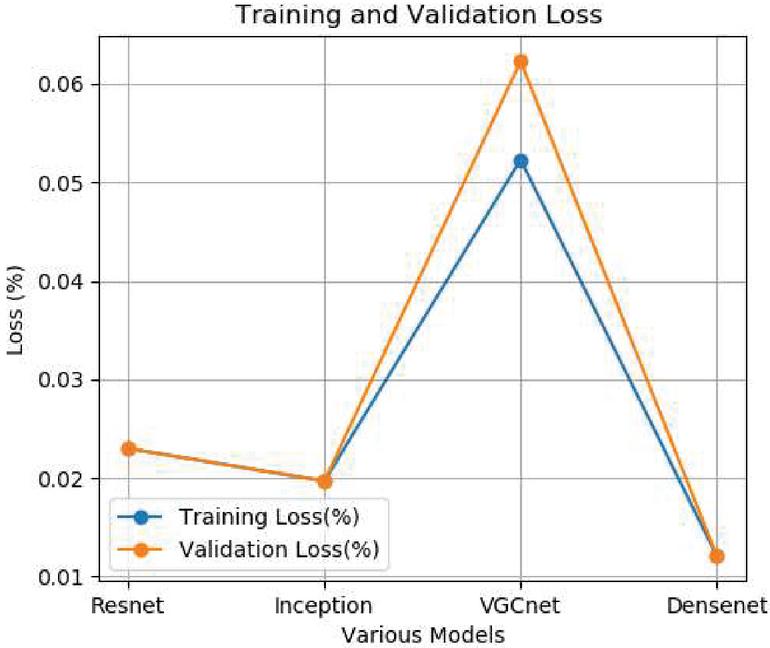
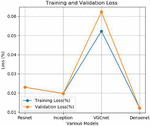
Figure 4 Training and validation loss.
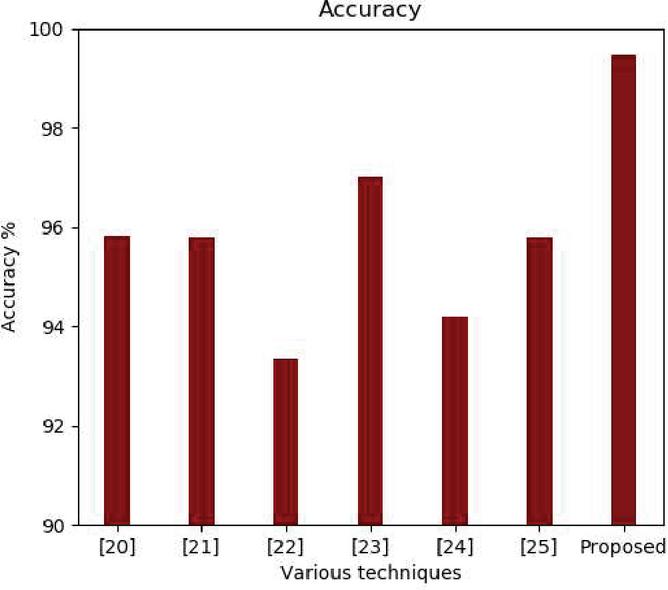
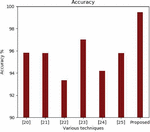
Figure 5 Accuracy comparison.
The training and validation loss is also experimented for all the four models. The average training loss for Resnet is 0.023%, Inception is 0.019%, VGCnet is 0.06% and Densenet is 0.010%. The results of the proposed method denotes that the densenet has higher accuracy and lower loss percentage compared to the remaining other models. The accuracy of the proposed approach is compared with the existing state-of-the-art approaches. The accuracy comparison of different approaches is shown in the Figure 5.
5 Conclusion and Future Enhancement
The major goal of the rice plant disease detection method is to provide quick and accurate disease predictions in crop disease control. Early detection of rice plant disease aids paddy scientists and producers in taking prompt actions to safeguard the plant. The proposed work evaluates the deep learning convolutional neural network models for rice plant disease identification. The architectures evaluated in this proposed approach are Inception, VGCnet, Resnet and Densnet [14, 15]. According to the results of the experiment, DenseNets seems to have a propensity to provide a consistent increase in accuracy, with no signs of performance degradation or overfitting. Furthermore, DenseNets requires a substantially lower number of parameters and a reasonable amount of computing time to achieve the top results in classification demonstrations. The average accuracy yield by the densenet is 99.45%. As a result, DenseNets is an excellent architecture for rice plant disease identification. In future the proposed approach can be enhanced further by reducing the computational time.
References
[1] Ferentinos, Konstantinos P. “Deep learning models for plant disease detection and diagnosis.” Computers and Electronics in Agriculture 145 (2018): 311–318.
[2] Ramcharan, Amanda, Kelsee Baranowski, Peter McCloskey, Babuali Ahmed, James Legg, and David P. Hughes. “Deep learning for image-based cassava disease detection.” Frontiers in plant science 8 (2017): 1852.
[3] Ashqar, Belal AM, and Samy S. Abu-Naser. “Image-based tomato leaves diseases detection using deep learning.” (2018).
[4] Arsenovic, Marko, Mirjana Karanovic, Srdjan Sladojevic, Andras Anderla, and Darko Stefanovic. “Solving current limitations of deep learning based approaches for plant disease detection.” Symmetry 11, no. 7 (2019): 939.
[5] Amara, Jihen, Bassem Bouaziz, and Alsayed Algergawy. “A deep learning-based approach for banana leaf diseases classification.” Datenbanksysteme für Business, Technologie und Web (BTW 2017)-Workshopband (2017).
[6] Jiang, Peng, Yuehan Chen, Bin Liu, Dongjian He, and Chunquan Liang. “Real-time detection of apple leaf diseases using deep learning approach based on improved convolutional neural networks.” IEEE Access 7 (2019): 59069–59080.
[7] Mohanty, S.P., Hughes, D.P., Salathé, M., 2016. Using deep learning for image-based plant disease detection. Front. Plant Sci. 7 (September), 1–7.
[8] Barbedo, Jayme GA. “Factors influencing the use of deep learning for plant disease recognition.” Biosystems engineering 172 (2018): 84–91.
[9] Rangarajan, Aravind Krishnaswamy, Raja Purushothaman, and Aniirudh Ramesh. “Tomato crop disease classification using pre-trained deep learning algorithm.” Procedia computer science 133 (2018): 1040–1047.
[10] R. Kaur and V. Kaur, “A deterministic approach for disease prediction in plants using deep learning,” 2018.
[11] Nagasubramanian, Koushik, Sarah Jones, Asheesh K. Singh, Arti Singh, Baskar Ganapathysubramanian, and Soumik Sarkar. “Explaining hyperspectral imaging based plant disease identification: 3D CNN and saliency maps.” arXiv preprint arXiv:1804.08831 (2018).
[12] Rajasekar, Vani, J. Premalatha, and K. Sathya. “Cancelable Iris template for secure authentication based on random projection and double random phase encoding.” Peer-to-Peer Networking and Applications 14, no. 2 (2021): 747–762.
[13] Suresh, G., V. Gnanaprakash, and R. Santhiya. “Performance Analysis of Different CNN Architecture with Different Optimisers for Plant Disease Classification.” In 2019 5th International Conference on Advanced Computing & Communication Systems (ICACCS), pp. 916–921. IEEE, 2019.
[14] Verma, Shradha, Anuradha Chug, Amit Prakash Singh, Shubham Sharma, and Puranjay Rajvanshi. “Deep learning-based mobile application for plant disease diagnosis: A proof of concept with a case study on tomato plant.” In Applications of Image Processing and Soft Computing Systems in Agriculture, pp. 242–271. IGI Global, 2019.
[15] Wu, Harvey, Tyr Wiesner-Hanks, Ethan L. Stewart, Chad DeChant, Nicholas Kaczmar, Michael A. Gore, Rebecca J. Nelson, and Hod Lipson. “Autonomous detection of plant disease symptoms directly from aerial imagery.” The Plant Phenome Journal 2, no. 1 (2019): 1–9.
[16] Rathore, N. P. S., and Prasad, L. (2020). Automatic rice plant disease recognition and identification using convolutional neural network. Journal of Critical Reviews, 7(15), 6076–6086.
[17] Sethy, P. K., Barpanda, N. K., Rath, A. K., and Behera, S. K. (2020). Nitrogen deficiency prediction of rice crop based on convolutional neural network. Journal of Ambient Intelligence and Humanized Computing, 11(11), 5703–5711.
[18] Moslehi, F., Haeri, A. An evolutionary computation-based approach for feature selection. J Ambient Intell Human Comput 11, 3757–3769 (2020). https://doi.org/10.1007/s12652-019-01570-1
[19] Matin, M. M. H., Khatun, A., Moazzam, M. G., and Uddin, M. S. (2020). An Efficient Disease Detection Technique of Rice Leaf Using AlexNet. Journal of Computer and Communications, 8(12), 49.
[20] Patidar, S., Pandey, A., Shirish, B. A., and Sriram, A. (2020, July). Rice plant disease detection and classification using deep residual learning. In International Conference on Machine Learning, Image Processing, Network Security and Data Sciences (pp. 278–293). Springer, Singapore.
[21] Xiao, M., Ma, Y., Feng, Z., Deng, Z., Hou, S., Shu, L. and Lu, Z. (2018) Rice Blast Recognition Based on Principal Component Analysis and Neural Network. Computers and Electronics in Agriculture, 154, 482–490. https://doi.org/10.1016/j.compag.2018.08.028
[22] Prajapati, H.B., Shah, J.P. and Dabhi, V.K. (2017) Detection and Classification of Rice Plant Diseases. Intelligent Decision Technologies, 11, 357–373. https://doi.org/10.3233/IDT-170301
[23] Ahmed, K., Shahidi, T.R., Irfanul Alam, S.M. and Momen, S. (2019) Rice Leaf Disease Detection Using Machine Learning Techniques. 2019 International Conference on Sustainable Technologies for Industry 4.0 (STI), Dhaka, 24–25 December, 1–5. https://doi.org/10.1109/STI47673.2019.9068096
[24] Islam, T., Sah, M., Baral, S. and Choudhury, R.R. (2018) A Faster Technique on Rice Disease Detection Using Image Processing of Affected Area in Agro-Field. 2018 Second International Conference on Inventive Communication and Computational Technologies (ICICCT), Coimbatore, 20–21 April 2018, 62–66. https://doi.org/10.1109/ICICCT.2018.8473322
[25] Maniyath, S.R., Vinod, P., Niveditha, M., Pooja, R., Shashank, N., Hebbar, R., et al., (2018) Plant Disease Detection Using Machine Learning. 2018 International Conference on Design Innovations for 3Cs Compute Communicate Control (ICDI3C), Bangalore, 25–28 April 2018, 41–45.
Biographies

S. Babu is a researcher at SRM Institute of Science and Technology, Kattankulathur. His current research focus is on Networking, Cloud Computing, and Software Engineering. He has completed Bachelor, Masters, and Ph.D. in Computer Science Engineering from Anna University, Chennai India.

M. Maravarman received his M.E in Computer and Communication, Anna University, Chennai. Currently, he is pursuing a Ph.D. in Computer Science and Engineering from SRM University, Chennai. His area of interest includes Networking and Cloud Computing.

R. Pitchai is an Associate Professor at the Department of Computer Science and Engineering from the B V Raju Institute of Technology. His current research focus is on Cloud Computing and securing Cloud environments. He holds his B.E. degree in Computer Science and Engineering from Anna University, India in 2005 and M.Tech in Software Engineering at Bharathidasan University, Trichy in 2008, and completed Ph.D. in Information and Communication Engineering at Anna University Chennai. He has published around 25 papers in various reputed journals.
Journal of Mobile Multimedia, Vol. 18_3, 757–770.
doi: 10.13052/jmm1550-4646.18314
© 2022 River Publishers