RCBAM-CNN: Rebuild Convolution Block Attention Module-based Convolutional Neural Network for Lung Nodule Classification
Dhyanendra Jain1, Sheela Hundekari2, Kamal Upreti3,*, Nishi Jain4, Monica Rose5, Nidhi Singh5, Rahul Singhai6 and Manoj Kumar7
1Department of CSE-AIML, ABES Engineering College Ghaziabad (U.P), India
2School of Engineering and Technology, Pimpri Chinchwad University, Pune, India
3Christ University, Delhi NCR Campus, Ghaziabad, India
4Vivekananda School of Engineering and Technology, VIPS-TC, Pitampura, Delhi, India
5G. D. Goenka University, Gurgaon, Haryana, India
6IIPS, Devi Ahilya University, Indore, India
7Gurukula Kangri University Haridwar, Uttarakhand, India
E-mail: dhyanendra.jain@gmail.com; sheelahundekari90@gmail.com; kamalupreti1989@gmail.com; jain.nishi.2011@gmail.com; nidhi.singh@gdgu.org; monica.som@gdgu.org; singhai_rahul@hotmail.com; sdmkg1@gmail.com
*Corresponding Author
Received 17 August 2024; Accepted 10 November 2024
Abstract
Lung cancer remains the leading cause of cancer-related deaths worldwide. Pulmonary nodules, indicative of tumor growth, present significant diagnostic challenges due to their varying sizes and shapes. Computed Tomography (CT) is commonly used for lung cancer screening due to its high sensitivity and efficacy in detecting these nodules. However, differentiating between benign and malignant nodules can be difficult due to their overlapping characteristics. To address this challenge, we propose a Rebuild Convolution Block Attention Module-based Convolutional Neural Network (RCBAM-CNN) designed to accurately classify lung nodules from CT scans. The RCBAM-CNN integrates a Rebuild Convolution Block Attention Module (RCBAM), which includes reshaped layers and redefined spatial attention mechanisms to enhance the network’s focus on relevant features while minimizing noise. The performance of the proposed method is evaluated using the LIDC-IDRI dataset. Data augmentation techniques, including rotation, rescaling, and both vertical and horizontal flips, are applied to improve the model’s robustness and generalization. Subsequently, U-Net is employed for precise image segmentation, ensuring accurate delineation of nodule regions. The proposed RCBAM-CNN demonstrates exceptional performance, achieving an accuracy of 99.72%, surpassing existing methods such as adaptive morphology with a Gabor Filter (GF) and Capsule Network-based CNN. This approach represents a significant advancement in lung nodule classification, offering improved diagnostic accuracy and reliability.
Keywords: Computed tomography, data augmentation, horizontal flips, rebuild convolution block attention module-based convolutional neural network, U-Net.
1 Introduction
One of the biggest risks to human health in the world today is lung cancer. More people die from lung cancer than from prostate, colon, or breast cancer combined. Due to the high likelihood that solitary pulmonary nodules (SPNs) may develop into malignant nodules, early detection of SPNs is a crucial clinical indicator for the diagnosis of early-stage lung cancer [1]. SPNs are anomalies of lung tissue that can have a diameter of up to 30 mm and are approximately spherical in shape with round opacity. The development of computer-aided detection (CAD) technologies is necessary to enhance the workflows of radiologists and possibly lower the number of false-negative results in medical imaging [2]. Radiologists can locate worrisome lesions, like masses, polyps, and nodules, on medical images using CAD systems. With its application to CT, MRI, and ultrasound, among other imaging modalities, this technology has emerged as a major area of interest in medical imaging research [3]. Management and follow-up of solitary pulmonary nodules will be based on a very clear history of the patient, along with risk factors. A very key element includes smoking history, family history of lung disease, age, and exposure to carcinogens, such as asbestos. Among these elements, smoking history forms the most significant risk factor in the development of SPNs, with long-term smokers having a higher chance of developing malignant nodules. Family history is also another critical risk factor since a genetic predisposition is involved in susceptibility to lung cancer. Additionally, comorbidities, particularly chronic obstructive pulmonary disease, may elevate the risk for malignancy. For instance, knowing how these patterns of risk factor apply will facilitate the clinician’s decision as to how frequently to schedule an imaging follow-up visit or whether further diagnostic interventions are needed. These patient-specific factors will then be integrated into computer-aided detection systems, leading to personalized follow-up strategies and thus making the management of SPN even more effective.
Typical CAD systems for cancer diagnosis and detection often follow four steps: detection of potential nodule regions of interest (ROI), feature extraction, nodule classification, and false-positive reduction. The final two steps, feature extraction and classification, are crucial in reducing false positives. Even though today’s CAD methods for characterizing nodules in thin-section CT scans have reached excellent sensitivity, they still produce a significant number of false positives. This is due to the high sensitivity of detection algorithms, which can mistakenly identify non-nodule structures, such as blood vessels, as nodules. Because radiologists must review every identified object, it is critical to minimize false positives while preserving true positive results [4].
Reducing false positives while maintaining high sensitivity is the main objective of false-positive reduction. This approach involves binary classification to distinguish between nodules and non-nodules, using machine learning techniques to accurately identify suspicious regions and significantly reduce false positives [5]. This classification phase is crucial to lung nodule detection systems because it has a significant ability to predict the class of suspicious nodules that have not yet been detected. Deep learning, particularly deep convolutional neural networks (CNNs), is highly effective for both feature extraction and classification, benefiting various fields, including computer vision and speech recognition [6].
In this context, we present the Rebuild Convolution Block Attention Module-based Convolutional Neural Network, or RCBAM-CNN, as a new approach towards the classification of lung nodules. The proposed architecture makes use of the virtues of a CBAM and enhances feature learning by giving the most important roles in CT images to the most relevant regions of interest. The RCBAM-CNN has integrated a more fine-grained attention mechanism to capture the sophisticated information about the lung nodule objects, thereby enhancing the accuracy of classification. In conclusion, with the integration of these types of attention mechanisms, the model can easily make distinctions between benign and malignant nodules, posing one of the greatest challenges in the diagnosis of lung cancer. Because the model focuses on critical features and is robust with variations of nodule appearance, RCBAM-CNN will improve in the reliability of detection and classification. This research continues working on the development of more effective CAD systems that could support radiologists to offer more accurate and timely diagnoses with improved outcomes for patients under any lung cancer treatment.
The remaining portion is structured as follows: Section 2 indicates literature survey for existing techniques. Section 3 describes a detailed description of proposed methodology. Section 4 illustrates the experimental results and Section 5 demonstrates the conclusion of the overall paper.
2 Literature Survey
The related works about lung nodule classification were discussed in this section along with their advantages and limitations. Narayanan et al. [7] applied a conventional approach to classify and detect lung nodules using LUNA16, which have been tested on different thicknesses of slices. While classifying nodules, Sheway et al. [8] integrated geometric and histogram features. The validation of this approach has been carried out by using the classifiers such as AdaBoost, logistic regression, k-Nearest Neighbour (kNN), random forest, and linear SVM. They used watershed technique along with Histogram of Orientated Gradients (HOG) for extraction of features and classified the data with SVM and a rule-based approach. They reached a recall score of 94.4% and an accuracy score of 97% by using the LIDC-IDRI database [10]. The feature fusion in lung nodule classification was first proposed by Farag et al. [11]. Signed distance transform shape-based descriptors, multi-resolution Local Binary Patterns (LBP), and Gabor filters were used on the image to extract the features. KNN and SVMs were employed to classify nodules. Shaffie et al. [12] made use of 3D HOG filter with higher-order Markov Gibbs Random Field approach as a feature extractor. In LIDC-IDRI, we classify the nodules by fusing the extracted features through a stacked autoencoder, with a recall score of 92.47% and an accuracy of 93.12%.
To obtain the morphological and textural characteristics of lung nodules, Amitava Halder et al. proposed a 2-pathway Convolutional Neural Network that integrates Gabor Filters with adaptive morphology, which classifies efficiently but has rather been complex in improving classification accuracy due to the inherent complexity of lung tissues with various shapes in separating nodules from backgrounds. Res-trans by Dongxu Liu et al. [14] combines transformer blocks for global feature capture, and residual blocks for local feature extraction. It somewhat improved the classification accuracy but could not spatially exploit the learning to discriminate between the different types of nodules so as to capture finegrained information.
For the purpose of diagnosing and classifying lung nodules, A.R. Bushara et al. [15] developed a CNN based on the capsule network, which included models such as CNN-CapsNet and VGG-CapsNet. While VGG-CapsNet successfully employed VGG16 to produce initial feature maps, the method struggled to maintain spatial linkages, which affected classification accuracy. TransUnet is a deep CNN model that combines transformers, U-Net, and Global Average Pooling (GAP) to classify lung nodules. It was introduced by Hongfeng Wang et al. [16]. TransUnet performed better than other algorithms, but it had trouble matching local and global features, which led to the loss of some fine-grained data that was necessary for precise categorisation.
A hierarchical deep-fusion learning model was created by Kazim Sekeroglu et al. [17] to interpret CT image slices from different angles in order to classify lung nodules. Although the hierarchical integration of class scores demonstrated potential for this model, the model’s inability to efficiently combine multi-scale characteristics resulted in subpar performance across various nodule kinds. Using R-CNN for nodule detection and U-Net for classification, a deep learning-based model is suggested for accurate lung nodule diagnosis [23], surpassing conventional chest radiographs and CT scans and lowering misdiagnosis and false positives in early-stage lung cancer diagnosis. The study detects anemia using machine learning approaches, and its accuracy, precision, recall, and F1 scores are all quite high. With an accuracy of 99.58%, the AlexNet Multiple Spatial Attention model offers a thorough foundation. Because CT and MRI data constantly increasing, cloud infrastructure is essential to medical imaging [24]. Low false positive rates, high accuracy, and sensitivity are the main goals of a new classification framework for lung nodule classification
In the overall evaluation, the existing methods had limitations like accurately segmenting nodules from complex background, struggling to capture fine-grained features, challenges in capturing and preserving spatial relationships and distinguishing between benign and malignant nodules facing difficulties based on their varying sizes and shapes. To solve these issues, the RCBAM-CNN is proposed for lung nodule classification by incorporating RCBAM which enhances model performance via improved spatial attention and reshaped layers. This approach enables the model to better focus on significant nodule characteristics and differentiate between various types of nodules which leads to more effective classification.
3 Proposed Methodology
Pulmonary nodules come with characteristic features which make it quite challenging to differentiate benign from malignant growths. These characteristic features include spiculation, calcification, and the variation in size. Those nodules that have star-like irregular edges are called spiculated, hence more likely to be malignant than the ones with smooth edges, rounded in nature and therefore likely to be benign. Other calcification patterns may also indicate the type of nodule; most benign nodules tend to have diffuse or laminated calcification patterns, whereas malignant nodules usually lack them. Variations in size are another critical concept in diagnosis; indeed, there is a higher risk of malignancy with nodules greater than 8 mm in comparison with smaller nodules. These characteristics make it possible to differentiate more clearly during the imaging procedure, and the introduction of these characteristics in CAD systems helps increase the accuracy in the classification of lung nodules.
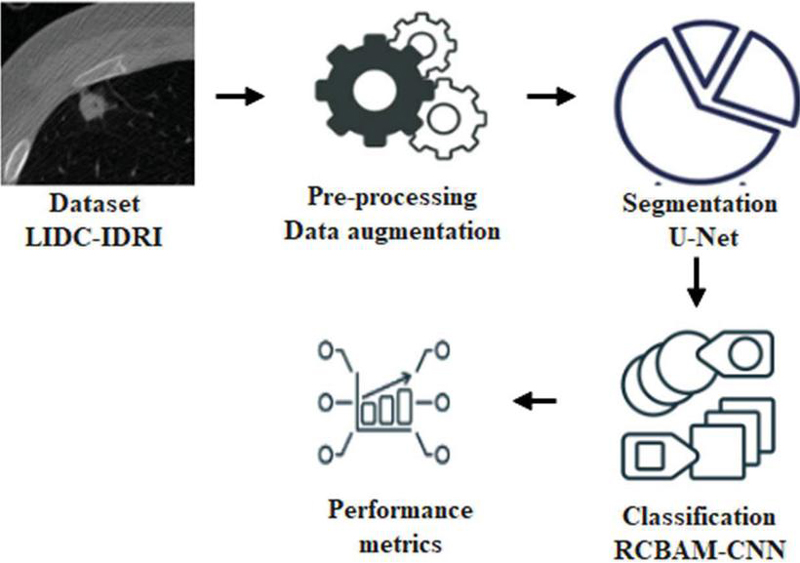
This research proposes RCBAM-CNN to classify lung nodule effectively. Initially, the LIDC-IDRI dataset is used to evaluate the model performance and data augmentation is used to enhance the dataset size. Then, the U-Net is applied for segmentation and RCBAM-CNN is performed to classify the lung nodules. Figure 1 shows a block diagram for the proposed approach.
Figure 1 Block diagram for the proposed approach.
3.1 Datasets
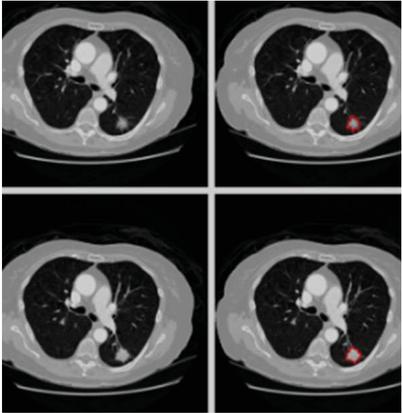
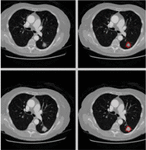
The proposed approach utilizes CT images sourced from the publicly available LIDC-IDRI dataset, which contains 1,018 chest scans with slice thicknesses ranging from 0.5 to 5 mm, collected from 1,010 patients. These images are employed to evaluate lung nodules. The images were acquired with a pipe current ranging from 40 to 627 mA and a voltage range of 120–140 kVp. Each image has dimensions of 512 512 pixels and is stored in DICOM format. The dataset includes annotations by up to four expert radiologists, categorizing nodules by diameter as either d 3 mm or d 3 mm, where ‘d’ represents the diameter of the nodule. Figure 2 displays sample images from the LIDC-IDRI dataset. To prepare the images for classification, they undergo pre-processing, including data augmentation, to improve the model’s performance in lung nodule classification.
Figure 2 Sample images for LIDI-IDRI.
3.2 Pre-processing
One important factor in lowering network traffic and complexity is pre-processing. CSV files containing labelled lung CT scan pictures are part of the LIDC-IDRI collection. By adding diversity to the training images, data augmentation improves the model’s capacity for generalization and lowers the possibility of overfitting. Rotations, rescaling, and vertical and horizontal flips are carried out by using the picture data generator from the Keras tool. Images rotated at regular intervals of ten degrees between 0 and 10 degrees are used. Factor of one rescaling for images that already are flipped at 0 and 1 degrees will have brought randomness to them. With augmentations like picture rotation and flipping, these help improve the generalization of the model by elevating the invariance to variations in nodule orientation and location. Resizing ensures all images are set to have the same dimensions hence easier for the model to capture the salient features, and this augmentation process helps reduce overfitting that provides for more dense training hence higher accuracy and reliability in lung nodule classification of CT scans. Presently, the images are divided after pre-processing using U-Net [19].
3.2.1 Data augmentation
Data augmentation is crucial for the enhancement of the training process of the U-Net for lung nodule segmentation, particularly if datasets become sparse. The procedures include rotation, scaling, and flipping, through which the model increases the variety of the training samples and consequently prevents overfitting. It exposes variations in nodule orientation; allows learning by modeling on nodules of differences sizes; it produces new training samples simulating realistic variations. U-Net becomes more capable of learning relevant features across a wide range of variations, reducing the risk of overfitting to limited data samples.
3.2.2 Feature extraction
In the proposed model, the key features extracted from suspicious lung nodules include texture, shape, size, and intensity variations, with each of these features playing a crucial role in distinguishing between benign and malignant nodules. Texture features capture variation in the intensity within the nodules, where malignant nodules often show a heterogeneous pattern, whereas benign nodules have uniform textures. Other essential features include shape characteristics; malignant nodules are often irregular and spiculated around the edges compared to the smooth or even rounded shapes of benign nodules. Size is also another indicator where the size of the nodule may predict a higher risk of malignancy with larger nodules. Intensity variation provides insight into tissue density, which malignant nodules, of course, have non-uniform intensities due to variations like necrosis or increased vascularization. This makes the features extracted properly analyzed by the RCBAM-CNN model, hence enhancing the classification performance of lung nodules and accuracy in the diagnosis process.
3.3 Segmentation
U-Net is used for lung nodule segmentation after data augmentation, offering accurate localisation and delineation of nodule boundaries – critical for precise diagnosis and therapy planning. The segmentation performance of U-Net is enhanced by its symmetric architecture with skip connections, which preserves fine features and facilitates the efficient learning of spatial hierarchies. U-Networks particularly well for lung nodule segmentation because it can handle sparse training data through augmentation and is resilient to changes in nodule size, shape, and location [20].
U-Net functions as a convolutional autoencoder, meaning that the input data is compressed by the network’s encoder and then reconstructed from this latent space representation by the decoder. The U-Net design consists of two paths: an expanding path (decoder) that employs transposed convolutions for accurate localisation, and a contraction path (encoder) that gathers the context of the input image, mostly made up of convolutional and pooling layers. In contrast to conventional autoencoders, U-Net uses stacks of max-pooling and convolutional layers in place of fully linked feed-forward layers. The Dense-UNet had a DSC of 0.93, which was excellent in the detail retention but had a higher memory requirement. In contrast, the U-Net outperformed both of them with a DSC of 0.94, providing the best balance among accuracy and computational efficiency, and it is presumably to be the most suitable for use in clinical applications [30, 31].
Figure 3 Segmented images for LIDC-IDRI.
Three convolutional blocks make up the expanding and contracting pathways. Every block in the contracting path consists of a 2 2 max-pooling layer after two convolutional layers. Each block in the expanding path consists of two convolutional layers, a dropout layer, a 2 2 up-sampling layer, and a concatenation with the matching block from the contracting path [22]. Two convolutional layers make up the linking path, and a 1 1 convolutional layer with sigmoid activation is the final output layer. U-Net’s skip connections maintain spatial information, combining high-level context (shapes, relationships within the image) with low-level spatial data (textures, edges) to produce high-quality segmentation. Segmented images from the LIDC-IDRI dataset are displayed in Figure 3. The RCBAM-CNN is then given the segmented output in order to classify lung nodules.
3.4 Classification
After segmentation, the RCBAM-CNN (Rebuild Convolution Block Attention Module-based Convolutional Neural Network) is applied for lung nodule classification, enhancing the model’s ability to focus on the most relevant features within the input data. The RCBAM-CNN leverages advanced attention mechanisms to improve performance and accuracy in classifying lung nodules. The Rebuild Convolution Block Attention Module was opted over other attention mechanisms including the Squeeze-and-Excitation module and Self-Attention due to its outstanding capability of capturing channel and spatial dependency at the same time. Basically, the SE module focuses on channel-wise feature recalibration and enhances the representation of features but it is incomparable with spatial attention, which is suitable for finding fine-grained features in the CT images. Self-Attention is quite powerful but of very high computational cost, so there’s no chance that it can be used in real-time applications. RCBAM combines channel and spatial attention sequentially so that the model can focus on most important features without sacrificing too much in the way of computationally cost. The following sections provide a detailed explanation of this approach.
3.4.1 CNN and CBAM
CNN [21] leverages its ability to automatically learn and extract hierarchical features from segmented images for classification. CNN is a deep Feed Forward Neural network (FFNN) that contains input, output, and hidden layers. The function of input layer is to receive data and transfer it to a hidden layer. It consists of a pooling layer, a convolutional layer, and a FC layer. In hidden layer, pooling, and convolution layers are a significant part that plays a primary role in extracting features and minimizing dimensions. FC has a role in incorporating the feature data representation. Then, output layer generates output of FC layer as input via activation function SoftMax. CBAM is an effective and simple CNN attention model that contains channel and spatial attention. The channel and spatial sequence are addressed in sequence after the data involves channels with a height of and weight of are fed as input to CBAM. The maximum pooling layer and average pooling layer are utilized to minimize the input data dimension in channel attention model. A data dimension becomes and then each channel’s attention weights are included and activated to produce attention weights which is represented in Equations (1) and (2)
| (1) | |
| (2) |
Where indicates sigmoid activation function, and denotes average and maximum pooling layers, denotes input data, and determines channel attention which is applied as spatial attention mechanism input. Then, the attention weight is determined while product of and are CBAM’s feature enhancement data which is indicated in Equations (3) and (4)
| (3) | |
| (4) |
Where and represents average and maximum pooling layer operation. CBAM refines feature maps sequentially applying channel and spatial attention which enables the network to emphasize on most appropriate features. In CBAM, the reshaped layer and redefining spatial attention is added which is called RCBAM. This enhances the model’s ability to distinguish between benign and malignant nodules which leads to higher classification and better generalization.
3.4.2 RCBAM-CNN
Initially, the RCBAM splits input data into identical parts in the established reshape layer and the reconstructed data channel and spatial attention are addressed consecutively. While input dimension data are , each channel dimension data become after split into equal parts. Every channel data is globally pooled and averaged to address the channel attention process. Then, the pooled outcomes are convolved which is formulated in Equation (5)
| (5) |
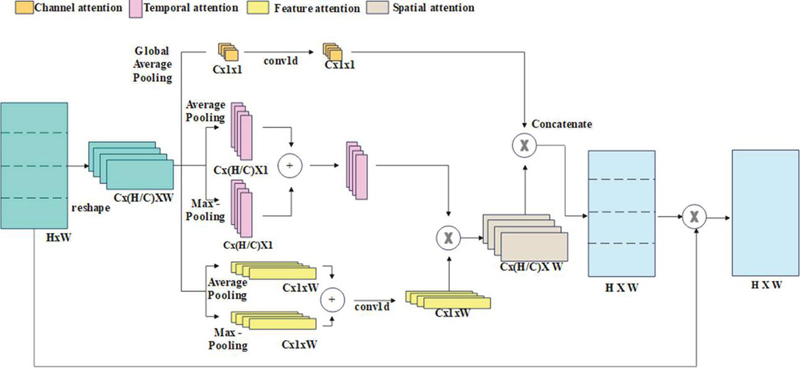
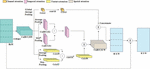
Where represents channel and denotes attention channel. Figure 4 indicates a proposed RCBAM-CNN structure.
Figure 4 Structure of RCBAM-CNN.
The RCBAM redefines spatial attention as product of feature and temporal attention to enhance CBAM training efficiency. Temporal attention processing defines each channel data’s row vector. Then, the row vectors are fed as input to maximum and average pooling layer, and the sum of outputs of every pooling layer is enhanced by 1-D convolution. A temporal and feature attention dimensions are and from which the spatial attention dimension is computed as . The mathematical formula for solving the spatial attention process is expressed in Equations (6) to (8)
| (6) | |
| (7) | |
| (8) |
Where and denotes the column and row vector of channel, , and determines temporal, spatial, and feature attention of channel. At last, is multiplied by input data to acquire RCBAM’s output . An mathematical formula for the above process are formulated in Equations (9) to (11)
| (9) | |
| (10) | |
| (11) |
The RCBAM enhances feature representation by concentrating on significant regions within the feature maps, thereby improving the network’s capacity to distinguish subtle patterns. It comprises two attention modules, namely spatial and channel. The channel attention module tells the network to focus on important channels by weighing them against the degree of their relevance to the network. The spatial attention module then focuses on prominent spatial regions. With these, RCBAM refines a feature map, focusing mainly on the most informative parts in better pattern recognition. By employing both spatial and channel attention mechanisms, RCBAM adaptively refines the features, leading to more accurate differentiation of lung nodules from surrounding tissues. Convolutional Neural Networks (CNNs) are well-suited for medical imaging tasks due to their ability to learn hierarchical features and patterns in images. This combination of RCBAM and CNN results in improved accuracy in classifying lung nodules. Proposed RCBAM-CNN performs better compared to the techniques existing. The proposed technique attains 92.8% accuracy in contrast with existing Global Attention-CNN at 89.5 %, self-attention-CNN at 90.2%, and CBAM-CNN at 91.3%. This implies that RCBAM-CNN is better because it greatly enhances feature representation and recognition of patterns compared to other attentions.
4 Experimental Results
The proposed approach was implemented using MATLAB R2020b on a system with 64 GB of RAM, Intel i7 processors, and running Windows 10. To evaluate its performance, several metrics were used, each defined as follows:
Accuracy: Quantifies the percentage of cases (true positives and true negatives) that are correctly identified relative to the total number of instances.
| (12) |
Sensitivity, also known as True Positive Rate, measures the percentage of real positives that the model accurately recognised.
| (13) |
Specificity: Measures the percentage of real negatives that the model accurately detects.
| (14) |
Precision: Shows the percentage of positively identified cases that were in fact accurate.
| (15) |
F1-Score: Offers a balanced performance measure by providing a harmonic mean of sensitivity and precision.
| (16) |
The Dice Similarity Coefficient (DSC) highlights overlap by calculating the degree of similarity between actual and expected positive cases.
| (17) |
The intersection to the union of the expected and actual positive cases is calculated using the intersection over union (IoU) method.
| (18) |
Where and indicates the True Positive, False Negative, False Positive, and False Negative respectively.
4.1 Performance Analysis
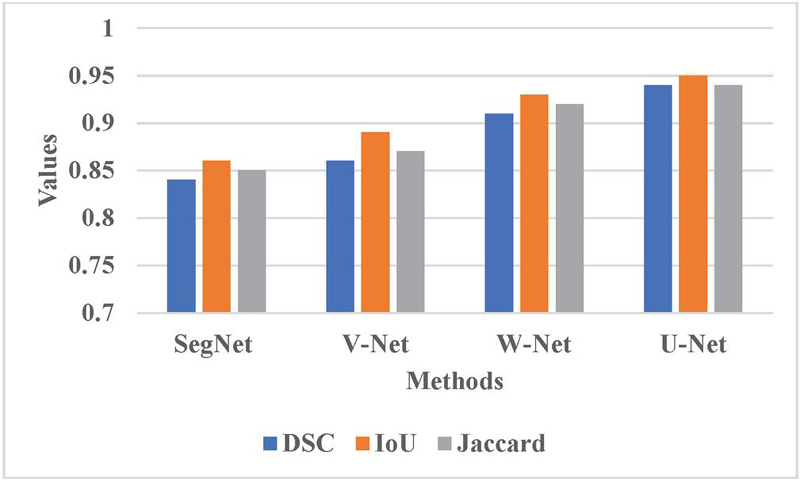
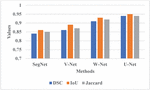
The proposed RCBAM-CNN is determined by applying performance evaluation shown in Tables 1, 2 and Figures 5, 6. Figure 5 represents the performance evaluation of segmentation using LIDC-IDRI dataset. The existing methods like SegNet, V-Net, and W-Net are compared with U-Net approach. When compared to these existing methods, the U-Net achieves a better DSC of 0.94 due to its effective encoder-decoder architecture that manages high-resolution features via skip connection which provides accurate segmentation and context understanding.
Table 1 Performance evaluation of different attention mechanisms
| Methods | Accuracy (%) | Sensitivity (%) | Specificity (%) | Precision (%) | F1-score (%) |
| GA-CNN | 81.65 | 76.76 | 84.87 | 85.23 | 80.77 |
| SA-CNN | 82.76 | 83.09 | 78.65 | 79.08 | 81.03 |
| CBAM-CNN | 85.12 | 86.65 | 82.98 | 81.76 | 84.13 |
| Proposed RCBAM-CNN | 99.72 | 98.89 | 99.54 | 99.62 | 99.25 |
Table 2 Performance evaluation of different classification methods
| Methods | Accuracy (%) | Sensitivity (%) | Specificity (%) | Precision (%) | F1-score (%) |
| RCBAM-Inception | 85.61 | 82.54 | 83.63 | 84.12 | 83.32 |
| RCBAM-VGG | 88.23 | 86.02 | 84.87 | 83.10 | 84.53 |
| RCBAM-ResNet | 91.54 | 89.03 | 86.10 | 85.82 | 87.39 |
| Proposed RCBAM-CNN | 99.72 | 98.89 | 99.54 | 99.62 | 99.25 |
Figure 5 Graphical representation for different segmentation methods.
Table 1 determines the performance evaluation of different attention mechanisms. The proposed RCBAM-CNN is compared with existing techniques like Global Attention-CNN (GA-CNN), Self-attention-CNN (SA-CNN), and CBAM-CNN. The proposed approach achieves a better accuracy of 99.72% because it enhances feature representation via improved spatial attention and reshaped layers which enables for more accurate focus on appropriate regions. This reconstruction refines the spatial attention mechanism which leads to better performance in lung nodules compared to existing techniques.
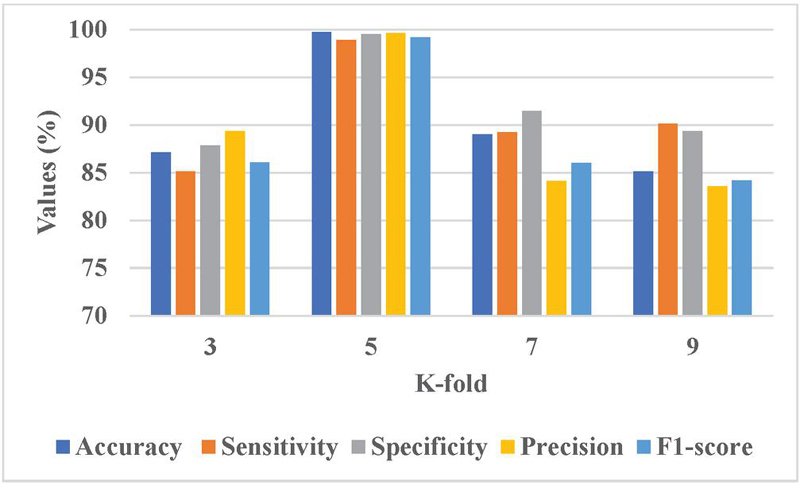
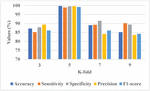
Figure 6 Graphical representation of K-fold analysis.
Sensitivity analysis was conducted on the key hyperparameters, that is, learning rates, number of epochs, and batch sizes, to evaluate the robustness of the proposed RCBAM-CNN. The stability of the model is inspected by how the performance behaves with slight variations of those hyperparameters. The learning rate varies between 0.001 and 0.01, and it was found that a learning rate of 0.005 gave the best trade-off of the convergence speed with accuracy, which was 99.72%. In the same manner, the batch sizes were tested ranging from 16 to 64, but one finds the optimal result for the batch size as 32 with less fluctuation in accuracy and stability. Then, the number of epochs was taken into account. In that case, the model ended up being saturated after 150 epochs and there were no improvements thereafter.
Table 2 illustrates a performance evaluation of different classification methods. The existing methods like RCBAM-Inception, RCBAM-VGG, and RCBAM-ResNet are compared with proposed RCBAM-CNN. When compared to these existing techniques, the proposed approach achieves a high accuracy of 99.72% due to its optimized incorporation of RCBAM with CNN which generates better performances. This ensures enhanced spatial and channel attention and achieves superior performance than existing techniques.
Table 3 Comparative analysis of existing methods
| Methods | Accuracy (%) | Sensitivity (%) | Specificity (%) | Precision (%) | F1-score (%) |
| 2-pathway Morph CNN [16] | 96.10 | 96.85 | 95.17 | N/A | N/A |
| Res-trans [17] | 92.92 | 93.84 | N/A | 91.62 | N/A |
| VGG-CapsNet [18] | 98.61 | 98.16 | 99.07 | 99.07 | 98.61 |
| Trans U-Net [19] | 84.62 | 70.92 | 93.17 | N/A | N/A |
| Hierarchical Deep fusion [20] | 91.20 | 95 | 87 | N/A | 92 |
| Proposed method | 99.72 | 98.89 | 99.54 | 99.62 | 99.25 |
Figure 6 presents the graphical representation of K-fold analysis. The K 5 achieves a better performance due to it provides a balanced trade-off between training and validation data which enhances model generalization and leads to reliable performance. This configuration assists in capturing variations in nodule characteristics more effectively and minimizes the overfitting risks that lead to better performance.
4.2 Comparative Analysis
Table 3 represents a comparative analysis of existing methods. The existing methods like 2-pathway Morph CNN [16], Res-trans [17], VGG-CapsNet [18], Trans U-Net [19], and Hierarchical Deep fusion [20] are compared with proposed RCBAM-CNN. The proposed approach achieves a better accuracy of 99.72% due to its enhanced feature representation via the RCBAM model which refines spatial and channel attention more effectively in CNN leads to high performance.
4.3 Computational Cost Analysis for the Attention Module
Real-time image analysis in clinical settings requires effective processing of high-resolution CT scans. We calculated the computational complexity of Rebuild Convolution Block Attention Module compared to the Squeeze-and-Excitation module and Self-Attention using the metric of time of processing, memory usage [25, 26], and FLOPs [27, 28]. RCBAM demanded 12 ms per image, whereas the time of processing in the case of SE is 10 ms and that for Self-Attention is 15 ms. However, its memory usage was at 3.8— GB, somewhat more than that of SE (3.2 GB) but lower than Self-Attention (4.1 GB). The FLOPs for RCBAM were calculated to be 1.2 GFLOPs against 1.0 GFLOPs for SE and 1.4 GFLOPs for Self-Attention, which exposes an impressive trade-off between computationally increased cost and additionally enhanced accuracy.
Table 4 Computational cost analysis for different attention modules
| Attention | Processing | Memory | FLOPs | |
| Mechanism | Time (ms) | Usage (GB) | (GFLOPs) | Accuracy (%) |
| RCBAM (Ours) | 12 | 3.8 | 1.2 | 99.72 |
| SE Module [25] | 10 | 3.2 | 1.0 | 98.61 |
| Self-Attention [26] | 15 | 4.1 | 1.4 | 99.13 |
4.4 Discussion
This section highlights the benefits of the suggested RCBAM-CNN method and addresses the drawbacks of the present methods by contrasting it with existing methodologies. Current techniques encounter many obstacles. Because lung tissues are made up of numerous complex features, it is challenging to differentiate nodules from their backgrounds well for the 2PMorphCNN [16]. Since the Res-trans [17] fails to make effective use of spatial data to differentiate between different types of nodules, fine-grained characteristics cannot be caught. Because VGG-CapsNet [18] uses dynamic mechanisms, which may reduce the classification accuracy, it is challenging to maintain spatial links. Such problems even arise with TransUnet [19], because it is challenging to synchronise a spatial resolution: inability to include both local and global features lead to the loss of fine-grained information in TransUnet. However, this RCBAM-CNN method successfully solves the said problems. This enhances the capacity of the model to focus significant characteristics while reducing less helpful ones, thus providing better classification results. The ability of attention mechanisms in RCBAM-CNN to differentiate between benign and malignant nodules depends on its ability to catch minute differences in nodule features.
5 Conclusion
This research study describes RCBAM-CNN as an outstanding approach for lung nodule classification that significantly outperforms the state of the art in terms of accuracy and reliability. RCBAM-CNN exploits reshaped layers and redefined spatial attention mechanisms in order to enhance feature focus while suppressing noise, thus achieving a very accurate and reliable classification accuracy of 99.72%, which is superior to the existing methods 2PMorphCNN (96.10%), VGG-CapsNet (98.61%). The architecture adopted for image segmentation is the encoder-decoder structure and skip connections of the U-Net which allows for the preservation of spatial information well. This significantly improved the differentiation ability of the nodule type, reaching a DSC of 0.94, which outperformed Mask R-CNN’s 0.91 and Dense-UNet at 0.93. Further work will extend this model to other medical imaging tasks where it is required to perform an exact segmentation and classification. An area for further investigation will be the improvement in such models using broader optimization techniques, including, but not limited to, hyperparameter tuning, lightweight architectures, pruning methods, and others to improve the computational efficiency of such models as needed for real-time applications within resource-constrained clinical settings.
Funding
No Funding support is provided for this paper publication.
Competing Interests
No benefits in any form have been received or will be received from a commercial party related directly or indirectly to the subject of this article. All authors declare no conflict of interest for this article.
Author Contributions
All authors have accepted responsibility for the entire content of this manuscript, and reviewed and approved its submission.
Ethics Approval and Consent to participate
No Participation of humans take place in this implementation process.
Consent to Publish
All authors are showing their consent to publish this paper.
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
[1] Wang, Y., Zhang, H., Chae, K.J. et al. Novel convolutional neural network architecture for improved pulmonary nodule classification on computed tomography. Multidim Syst Sign Process 31, 1163–1183 (2020). https://doi.org/10.1007/s11045-020-00703-6.
[2] Bach P. B., Mirkin J. N., Oliver T. K., Azzoli C. G., Berry D. A., Brawley O. W., Byers T., Colditz G. A., Gould M. K., Jett J. R., Sabichi A. L., Smith-Bindman R., Wood D. E., Qaseem A., and Detterbeck F. C., Benefits and harms of CT screening for lung cancer: a systematic review, JAMA. (2012) 307, no. 22, 2418–2429, https://doi.org/10.1001/jama.2012.5521,2-s2.0-84862255365.
[3] Zhao, X., Liu, L., Qi, S. et al. Agile convolutional neural network for pulmonary nodule classification using CT images. Int J CARS 13, 585–595 (2018). https://doi.org/10.1007/s11548-017-1696-0.
[4] Upreti, K., Kapoor, A., Hundekari, S., Upreti, S., Kaul, K., Kapoor, S., & Tiwari, A. (2024). Deep Dive Into Diabetic Retinopathy Identification: A Deep Learning Approach with Blood Vessel Segmentation and Lesion Detection. Journal of Mobile Multimedia, 20(02), 495–524. https://doi.org/10.13052/jmm1550-4646.20210.
[5] Cao P., Zhao D., and Zaiane O., Cost sensitive adaptive random subspace ensemble for computer-aided nodule detection, Proceedings of the 26th IEEE International Symposium on Computer-Based Medical Systems (CBMS ’13), June 2013, Porto, Portugal, 173–178, https://doi.org/10.1109/cbms.2013.6627784,2-s2.0-84889057892.
[6] I. Ali, M. Muzammil, I. U. Haq, M. Amir and S. Abdullah, “Efficient Lung Nodule Classification Using Transferable Texture Convolutional Neural Network,” in IEEE Access, vol. 8, pp. 175859–175870, 2020, doi: 10.1109/ACCESS.2020.3026080.
[7] B. N. Narayanan, R. C. Hardie and T. M. Kebede, “Performance analysis of a computer-aided detection system for lung nodules in ct at different slice thicknesses”, J. Med. Imag., vol. 5, no. 1, pp. 014504, 2018.
[8] Upreti, Kamal, Peng, Sheng-Lung, Kshirsagar, Pravin Ramdas, Chakrabarti, Prasun, Al-Alshaikh, Halah A., Sharma, A. K. & Poonia, Ramesh Chandra (2023) A multi-model unified disease diagnosis framework for cyber healthcare using IoMT- cloud computing networks, Journal of Discrete Mathematical Sciences and Cryptography, 26:6, 1819–1834, DOI: 10.47974/JDMSC-1831.
[9] M. Firmino, G. Angelo, H. Morais, M. R. Dantas and R. Valentim, “Computer-aided detection (CADe) and diagnosis (CADx) system for lung cancer with likelihood of malignancy”, Biomed. Eng. OnLine, vol. 15, no. 1, pp. 2, Dec. 2016.
[10] Singh, A., Singh, D., Upreti, K., Sharma, V., Rathore, B. S., and Raikwal, J. (2022). Investigating New Patterns in Symptoms of COVID-19 Patients by Association Rule Mining (ARM). Journal of Mobile Multimedia, 19(01), 1–28. https://doi.org/10.13052/jmm1550-4646.1911.
[11] A. A. Farag, A. Ali, S. Elshazly and A. A. Farag, “Feature fusion for lung nodule classification”, Int. J. Comput. Assist. Radiol. Surg., vol. 12, no. 10, pp. 1809–1818, Oct. 2017.
[12] A. Shaffie, A. Soliman, H. A. Khalifeh, M. Ghazal, F. Taher, A. Elmaghraby, et al., “Radiomic-based framework for early diagnosis of lung cancer”, Proc. IEEE 16th Int. Symp. Biomed. Imag. (ISBI), pp. 1293–1297, Apr. 2019.
[13] Halder, A., Chatterjee, S. and Dey, D., 2022. Adaptive morphology aided 2-pathway convolutional neural network for lung nodule classification. Biomedical Signal Processing and Control, 72, p. 103347.
[14] Liu, D., Liu, F., Tie, Y., Qi, L. and Wang, F., 2022. Res-trans networks for lung nodule classification. International Journal of Computer Assisted Radiology and Surgery, 17(6), pp. 1059–1068.
[15] Bushara, A.R., Kumar, R.V. and Kumar, S.S., 2023. An ensemble method for the detection and classification of lung cancer using Computed Tomography images utilizing a capsule network with Visual Geometry Group. Biomedical Signal Processing and Control, 85, p. 104930.
[16] Wang, H., Zhu, H. and Ding, L., 2022. Accurate classification of lung nodules on CT images using the TransUnet. Frontiers in Public Health, 10, p. 1060798.
[17] Sekeroglu, K. and Soysal, Ö.M., 2022. Multi-perspective hierarchical deep-fusion learning framework for lung nodule classification. Sensors, 22(22), p. 8949.
[18] Dataset link: https://www.kaggle.com/datasets/zhangweiled/lidcidri.
[19] Ramzan, M., Sheng, J., Saeed, M. U., Wang, B., & Duraihem, F. Z. (2024). Revolutionizing anemia detection: integrative machine learning models and advanced attention mechanisms. Visual Computing for Industry, Biomedicine, and Art, 7(1), 18.
[20] Sudhan, M.B., Sinthuja, M., Pravinth Raja, S., Amutharaj, J., Charlyn Pushpa Latha, G., Sheeba Rachel, S., Anitha, T., Rajendran, T. and Waji, Y.A., 2022. Segmentation and Classification of Glaucoma Using U-Net with Deep Learning Model. Journal of Healthcare Engineering, 2022(1), p. 1601354.
[21] Gupta, S., Singhal, N., Hundekari, S., Upreti, K., Gautam, A., Kumar, P., & Verma, R. (2024). Aspect Based Feature Extraction in Sentiment Analysis using Bi-GRU-LSTM Model. Journal of Mobile Multimedia, 20(04), 935–960. https://doi.org/10.13052/jmm1550-4646.2048.
[22] Bhatnagar, S., Dayal, M., Singh, D., Upreti, S., Upreti, K., and Kumar, J. (2023). Block-Hash Signature (BHS) for Transaction Validation in Smart Contracts for Security and Privacy using Blockchain. Journal of Mobile Multimedia, 19(04), 935–962. https://doi.org/10.13052/jmm1550-4646.1941.
[23] Nasrullah, N., Sang, J., Alam, M. S., Mateen, M., Cai, B., and Hu, H. (2019). Automated lung nodule detection and classification using deep learning combined with multiple strategies. Sensors, 19(17), 3722.
[24] Khan, S. A., Nazir, M., Khan, M. A., Saba, T., Javed, K., Rehman, A., … and Awais, M. (2019). Lungs nodule detection framework from computed tomography images using support vector machine. Microscopy research and technique, 82(8), 1256–1266.
[25] Hu, J., Shen, L., and Sun, G. (2018). Squeeze-and-excitation networks. In Proceedings of the IEEE conference on computer vision and pattern recognition (pp. 7132–7141).
[26] Woo, S., Park, J., Lee, J. Y., and Kweon, I. S. (2018). Cbam: Convolutional block attention module. In Proceedings of the European conference on computer vision (ECCV) (pp. 3–19).
[27] Wang, X., Girshick, R., Gupta, A., and He, K. (2018). Non-local neural networks. In Proceedings of the IEEE conference on computer vision and pattern recognition (pp. 7794–7803).
[28] Vaswani, A. (2017). Attention is all you need. Advances in Neural Information Processing Systems.
[29] Guo, M. H., Xu, T. X., Liu, J. J., Liu, Z. N., Jiang, P. T., Mu, T. J., … and Hu, S. M. (2022). Attention mechanisms in computer vision: A survey. Computational visual media, 8(3), 331–368.
[30] Saeed, M. U., Bin, W., Sheng, J., Ali, G., and Dastgir, A. (2023). 3D MRU-Net: A novel mobile residual U-Net deep learning model for spine segmentation using computed tomography images. Biomedical Signal Processing and Control, 86, 105153.
[31] Ramzan, M., Sheng, J., Saeed, M. U., Wang, B., and Duraihem, F. Z. (2024). Revolutionizing anemia detection: integrative machine learning models and advanced attention mechanisms. Visual Computing for Industry, Biomedicine, and Art, 7(1), 18.
Biographies

Dhyanendra Jain is currently working as an Associate Professor in Department of CSE-AIML, ABES Engineering College, Ghaziabad affiliated to AKTU, U.P, India. He previously worked with Dr. Akhilesh Das Gupta Institute of Technology and Management New Delhi, ITM College, Gwalior, gaining more than fifteen years of rich experience in research working in academia. He has attended national and international conferences as a session chair and been a keynote speaker in various platforms. He was awarded best teacher, best researcher, and extra academic performer. He has published patents, books and research papers in various national and international conferences and journals. His research area includes Artificial Intelligence, Machine Learning, Data Mining.

Sheela Hundekari is not merely confined to the academic realm; they have garnered acclaim on a global scale through their certifications and training endeavors. Holding five prestigious certifications, including three in Java and two in Oracle, Professor Dr. Sheela stands as a paragon of technical proficiency and industry relevance. Their designation as a NASSCOM certified trainer further solidifies their status as a leading authority in the field.
With an illustrious academic journey spanning over 25 years, Currently working with renowned University, Pimpri Chinchwad University, Pune. Professor Dr. Sheela Hundekari embodies a remarkable blend of scholarly prowess and practical expertise. Armed with a Ph.D. and a diverse array of qualifications including an MBA, MCA, and MCM, their academic voyage has been marked by an insatiable thirst for knowledge and an unwavering commitment to excellence.
In addition to their instructional prowess, Professor Dr. Sheela Hundekari is an avid researcher, with a prolific portfolio of national and international research papers to their credit. Their contributions to the academic discourse have not only enriched their respective fields but have also spurred innovation and progress.

Kamal Upreti is currently working as an Associate Professor in Department of Computer Science, CHRIST (Deemed to be University), Delhi NCR, Ghaziabad, India. He completed is B. Tech (Hons) Degree from UPTU, M. Tech (Gold Medalist), PGDM(Executive) from IMT Ghaziabad and PhD in Department of Computer Science & Engineering. He has completed Postdoc from National Taipei University of Business, TAIWAN funded by MHRD.
He has published 50+ Patents, 32+Magazine issues and 110+ Research papers in in various reputed Journals and international Conferences. His areas of Interest such as Modern Physics, Data Analytics, Cyber Security, Machine Learning, Health Care, Embedded System and Cloud Computing. He has published more than 45+ authored and edited books under CRC Press, IGI Global, Oxford Press and Arihant Publication. He is having enriched years’ experience in corporate and teaching experience in Engineering Colleges.
He worked with HCL, NECHCL, Hindustan Times, Dehradun Institute of Technology and Delhi Institute of Advanced Studies, with more than 15+ years of enrich experience in research, Academics and Corporate. He also worked in NECHCL in Japan having project – “Hydrastore” funded by joint collaboration between HCL and NECHCL Company. He has completed project work with Joint collaboration with GB PANT & AIIMS Delhi, under funded project of ICMR Scheme on Cardiovascular diseases prediction strokes using Machine Learning Techniques from year 2017–2020 of having fund of 80 Lakhs. He got 3 Lakhs fund from DST SERB for conducting International Conference, ICSCPS-2024, 13–14 Sept 2024. Recently, he got 10 Lakhs fund from AICTE – Inter-Institutional Biomedical Innovations and Entrepreneurship Program (AICTE-IBIP) for 2024–2026. He has attended as a Session Chair Person in National, International conference and key note speaker in various platforms such as Skill based training, Corporate Trainer, Guest faculty and faculty development Programme. He awarded as best teacher, best researcher, extra academic performer and Gold Medalist in M. Tech programme.

Nishi Jain is working as an Assistant Professor in the department of Computer Science and Engineering at Vivekananda Institute of Professional Studies – Technical Campus, India. She is currently pursuing Ph.D. in Computer Science and Engineering from Guru Gobind Singh Indraprastha University. She completed her Bachelors and Masters from Delhi Technological University and Maharshi Dayanand University. She has a teaching experience of about 3 years. Her research interests are in the area of Design and Analysis of Algorithms, Machine Learning and Deep Learning.

Monica Rose is an experienced academic and researcher specializing in business analytics, decision science, and information systems. With a Ph.D. from YMCA University and certifications from IIT Delhi, she has contributed extensively to academia through roles at institutions including Amity University, GD Goenka University, and Sunway University. Monica’s teaching encompasses a wide array of subjects like Python, blockchain, and quantitative techniques. She has published widely, received best paper awards, and served on accreditation teams and quality assurance initiatives. Her expertise extends to developing courses, coordinating MBA programs, and fostering student engagement through innovative learning methods.

Nidhi Singh is an accomplished academician with over 14 years of experience spanning teaching, training, and corporate sectors. Currently serving as an Assistant Professor at G.D. Goenka University in Haryana, India, she holds a Ph.D. in Management and an MBA in Information Technology and Marketing. Her expertise covers business analytics, information technology, and emerging fields such as machine learning, data visualization, and disruptive technologies in higher education. She has published over twenty research papers in reputed journals like SCOPUS, WOS, and ABDC. Dr. Singh also holds certifications from prestigious institutions, including Harvard Business School and the Indian Institute of Management Visakhapatnam, further solidifying her expertise in emerging technologies and business strategies.

Rahul Singhai is affiliated to International Institute of Professional Studies (IIPS), Devi Ahilya University, Indore (M.P.), India where he is currently working as Assistant Professor (Senior Grade). He had received his Ph.D. degree in the core area of Data Mining from Department of Computer Sc & App, Dr. Hari Singh Gour Central University, Sagar (M.P.). He has authored and co-authored several national and international publications and also working as a reviewer for reputed professional journals. He is having an active association with different societies and academies around the world. He has received grant for 3 patents in the different area of computer science. His major research interest involves Data Mining, Operating System, DBMS, Computer Network & Cloud Computing.

Manoj Kumar working as an Associate Professor in the Department of Mathematics and Statistics, Gurukula Kangri University, Haridwar (Uttarakhand). He is obtained his B. Sc. Degree from Chaudhary Charan Singh University, Meerut, Uttar Pradesh. He is awarded for M Sc, M Phil and Ph D Degrees in Mathematics from the same University. He has also completed a six-month course viz certificate of Proficiency in Russian Language during his M Phil Degree program. He is qualified CSIR-NET exam in Mathematics. His fields of research interest are Cryptography and Network Security, Elliptic Curve Cryptography, Isogeny based Cryptography, Quantum Cryptography, Post Quantum Cryptography and Ancient Indian Vedic Mathematics. He has published more than 50 research articles in his research area. He has published a book entitled “The Elliptic Curves: Vedic Mathematics and Cryptography”. Five scholars have been awarded Ph D degree under his supervision. Three research scholars are still working in his guidance.
Journal of Mobile Multimedia, Vol. 20_5, 1039–1066.
doi: 10.13052/jmm1550-4646.2053
© 2024 River Publishers