Learning by Experiencing: An Immersive Digital Twin Tool for ECG Education*
Daniel Flores-Martin1,*, Francisco Díaz-Barrancas2, Pedro J. Pardo2, Javier Berrocal2 and Juan M. Murillo2
1COMPUTAEX. Extremadura Supercomputing Center, Cáceres, Spain
2University of Extremadura, Badajoz, Spain
E-mail: daniel.flores@computaex.es; frdiaz@unex.es; pjpardo@unex.es; jberolm@unex.es; juanmamu@unex.es
*Corresponding Author
Received 30 October 2025; Accepted 15 December 2025
Abstract
Practical training in electrocardiogram (ECG) interpretation remains uneven, particularly in resource-limited settings, despite the central role of ECGs in cardiovascular diagnosis. This work evaluates whether an ECG-focused digital twin that integrates interactive simulation and deep learning guidance can achieve educationally valid realism, improve recognition of patterns and abnormalities through interactivity, and enhance accuracy and learner motivation via predictive feedback. We present ECGTwinMentor, a cross-platform system that synthesizes parameterized ECG waveforms, enables fine-grained control of physiologic variables, and delivers immediate predictive feedback for formative assessment. The diagnostic model supports low-latency inference on modest hardware. Validation with healthcare experts and medical students showed positive evaluations for realism, usability, and integration potential. Experts reported average ratings between 3.5 and 4.5 out of 5, while students rated usability between 4.6 and 4.8 and motivation and realism at 5.0, with most items scoring at least 4. These findings support the conclusion that an interactive, predictive digital twin can narrow the gap between theory and practice in ECG interpretation, offering an accessible, scalable, and reproducible approach to ECG education.
Keywords: Electrocardiogram, digital twins, edge computing, simulation, deep learning, formation.
1 Introduction
Cardiology remains a critical area of medicine due to the global impact of cardiovascular diseases, which continue to be the leading cause of mortality worldwide. Although advances in research and technology offer new opportunities to improve care, medical trainees often lack the necessary skills to interpret ECGs effectively, highlighting the need for better educational approaches [14].
Innovations in medical technologies, such as diagnostic tools and treatment techniques, have significantly enhanced the understanding and management of heart-related conditions. The application of cutting-edge technologies has led to more precise diagnostics, better patient outcomes, and increasingly personalized treatments, which ultimately improve public health and the quality of life for millions of people [26]. Also, artificial intelligence (AI) enhances ECG interpretation by identifying subtle patterns in large datasets, enabling early detection of cardiac and non-cardiac conditions, even in asymptomatic patients [24]. There are many fundamentals of supervised AI models and key applications, such as identifying left ventricular dysfunction, silent atrial fibrillation, and structural heart diseases [25]. Despite rapid advancements, there is a lack of relevant resources for personalized learning and a disconnect between AI technologies and their application in medical education [6].
Education and training in cardiology are crucial for healthcare students, as a solid understanding of cardiovascular diseases is vital for accurate diagnosis and treatment. Traditional methods, such as lectures, textbooks, and clinical rotations, provide foundational knowledge [20], but face limitations such as limited access to diverse ECG data, insufficient hands-on practice, and restricted exposure to various pathologies. Clinical settings can also be stressful and time-constrained. During the COVID-19 pandemic, many students were unable to complete rotations due to restricted hospital access, reducing opportunities for practical learning [11].
In recent years, digital twins have emerged as an innovative tool with the potential to revolutionize education across various disciplines, including healthcare [1]. A digital twin is a virtual representation of a physical system that simulates real-world conditions to support interactive learning and experimentation. In healthcare, digital twins have garnered attention for modeling complex medical systems and facilitating immersive, hands-on training, with applications in cardiology ranging from virtual heart models to simulated cardiac interventions [7]. These innovations enhance cardiology education by enabling students to practice ECG interpretation in an interactive, risk-free environment that bridges theoretical instruction and clinical application through realistic simulation and immediate feedback.
This paper presents ECGTwinMentor, an interactive digital twin platform that enhances cardiology education by integrating ECG simulation, DL-based diagnosis, and real-time visualization. Its main contributions include a predictive DL model trained on simulated ECG data, a web-based interface for interpretation, and a lightweight simulator deployable on both cloud and edge devices. The system’s novelty lies in its unified, modular design that supports scalable, hands-on learning across diverse environments.
This work builds on the hypothesis that a digital twin for ECG simulation and diagnosis can improve students’ accuracy and understanding in ECG interpretation compared to traditional resources. Accordingly, we explore the following research questions (RQ) with the associated hypothesis (H):
• RQ1: Can simulated ECGs achieve sufficient realism for educational use?
H1: The ECGTwinMentor simulator can reproduce ECG waveforms with sufficient realism to be valid for educational purposes.
• RQ2: Does interactive use of the twin enhance recognition of patterns and abnormalities?
H2: Interactive engagement with the simulator improves students’ ability to recognize ECG patterns and abnormalities compared to static resources.
• RQ3: How does predictive feedback influence learning outcomes and student engagement?
H3: Automated predictive feedback enhances both the accuracy of ECG interpretation and the motivation of learners.
The paper is organized as follows. Section 2 presents background and motivation; Section 3 describes the ECGTwinMentor system (DL model, web app, and simulator); Section 4 details the conducted experimental setup, while Section 5 reports expert and student feedback; Section 6 compares related approaches; Section 7 discusses findings; and Section 8 summarizes implications and future work.
2 Background and Motivations
Medical education in cardiology continues to face critical challenges. Limited access to diverse clinical ECG data, coupled with the intrinsic complexity of interpreting varied cardiac conditions, often leaves students and trainees underprepared to manage real-world cases [5]. Traditional teaching methods, such as lectures, textbooks, and occasional clinical exposure, offer only restricted opportunities for repetitive practice, controlled difficulty progression, and immediate feedback, which are essential for mastering ECG interpretation. This lack of experiential learning opportunities makes it difficult to develop robust skills within conventional training pathways.
In response to these challenges, technological advances have introduced digital twins and simulation-based learning as promising solutions. Cardiac digital twins can be generated from 12-lead ECGs, creating anatomically and physiologically realistic models that enable personalized analysis and training [12, 22]. At the population level, large-scale projects, such as those leveraging the UK Biobank, demonstrate the feasibility of constructing thousands of personalized digital twins, revealing key sex- and age-related variations in cardiac electrophysiology and showing how these tools can enrich both research and education [19]. These efforts highlight the potential of digital twins not only for personalized medicine but also for improving medical education by providing diverse, clinically grounded scenarios.
Simulation-based education has proven highly effective in medicine by providing safe, interactive environments for repeated diagnostic practice and feedback. Prior work shows that VR-based ECG learning improves interpretation accuracy, knowledge retention, and learner engagement [9], while virtual patient simulations enhance clinical reasoning beyond traditional didactic approaches. Although high-fidelity physical simulators remain valuable for cardiology training, they are resource-intensive and offer limited scalability and adaptability across learning contexts [8].
Within this evolving educational landscape, ECGTwinMentor integrates digital twin modeling and immersive simulation into a scalable platform for ECG education. By combining simulated ECG generation with AI-driven feedback, the system enables experiential, feedback-rich learning, while flexible deployment across centralized and edge environments supports low-latency, privacy-aware use in diverse educational contexts, effectively bridging conventional cardiology education with next-generation learning.
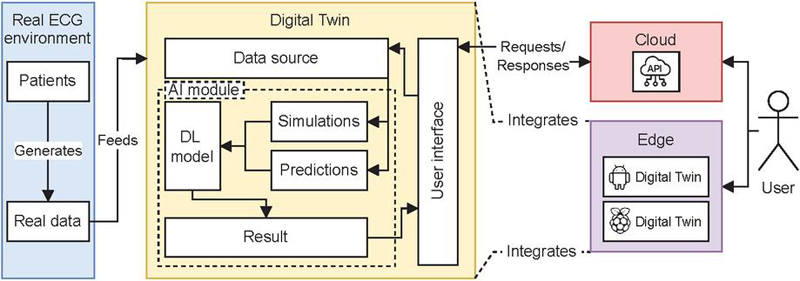
Figure 1 ECGTwinMentor system architecture.
3 ECGTwinMentor: System Overview
ECGTwinMentor comprises three core components: (1) a deep learning model for ECG-based cardiopathy prediction, (2) an interactive application providing diagnostic feedback across web, graphical, and edge deployments, and (3) a parameter-driven ECG simulator for exploring waveform variations and diagnostic patterns. These components are integrated into a unified digital twin platform that supports experiential learning and bridges theoretical instruction with practice, as illustrated in Figure 1.
• Real ECG environment: represents the physical environment where the digital twin aims to replicate real patients and their ECG information.
• Digital twin: the digital representation of the real-world ECG environment. It is composed of the following components:
– User interface: allows users to interact with the digital twin.
– Data source: provides the input data for the AI-based algorithm.
– AI module: responsible for processing the input data. It includes:
* Simulations: enables users to generate ECG simulations.
* Predictions: provides diagnostic predictions based on the user-inputs.
* DL model: processes the input and producing an output.
* Result: the output or diagnostic result returned to the user.
• Cloud: the digital twin can be accessed via cloud infrastructure through a custom web application and REST API.
• Edge: the digital twin can also be deployed on edge devices, allowing local execution and offline operation without the need for internet connectivity.
Although ECGTwinMentor is currently deployed in a cloud-based environment, its architecture supports both cloud and edge execution. Latency-sensitive components, such as ECG generation and visualization, are suitable for edge deployment, while computationally intensive tasks like model training and large-scale analytics are better handled in the cloud.
3.1 Deep Learning Model for Cardiopathy Prediction
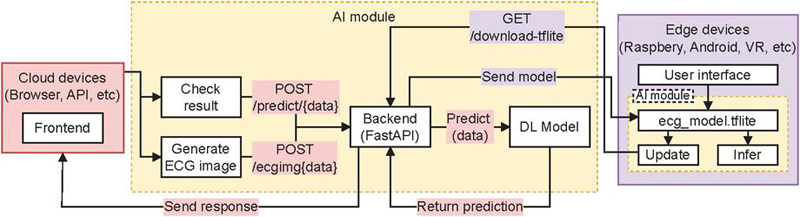
The core of ECGTwinMentor is the AI module (Figure 2). In cloud environments, it generates predictions, ECG images, and exports a lightweight model for edge use. In edge deployments, the module runs locally to perform inference and update the ecg_model.tflite, enabling offline operation and enhanced data security by keeping user data on the device.
Figure 2 ECGTwinMentor AI module.
3.1.1 Key ECG parameters
Evaluating ECG parameters in isolation may obscure relevant findings or lead to misinterpretation [21, 2]. These parameters describe the timing and morphology of ECG waves and intervals and are essential for cardiac diagnosis. Table 1 summarizes typical values, normal ranges, and common alterations associated with cardiopathies [16], including HR, rhythm, PR interval, QRS duration, ST segment, T wave, and QTc.
Table 1 ECG parameters with normal values and pathological variations
| HR | PR | QRS | QTc | |||
| (bpm) | Rhythm | (ms) | (ms) | ST Segment | T Wave | (ms) |
| 75 | Sinus | 160 | 90 | Normal | Normal | 400 |
| 45 | Sinus bradycardia | 180 | 100 | Normal | Normal | 420 |
| 110 | Sinus tachycardia | 140 | 80 | Normal | Normal | 390 |
| 90 | Atrial fibrillation | 180 | 110 | ST depression | Inverted | 450 |
| 85 | Sinus | 150 | 95 | Elevated (2 mm) | Normal | 410 |
| 72 | 1st degree AV block | 220 | 100 | Normal | Normal | 420 |
| 80 | Sinus | 160 | 130 | Normal | Normal | 390 |
| 60 | Sinus | 140 | 90 | Normal | Normal | 400 |
| 95 | Ventricular tachycardia | – | 160 | ST depression | Peaked | 500 |
| 55 | Sinus | 170 | 110 | Normal | Flattened | 430 |
These parameters are used to illustrate how variations in key electrocardiographic parameters correspond to specific cardiac conditions. Each row reflects a distinct ECG profile with plausible values derived from clinical norms and known pathological deviations.
3.1.2 Dataset and data generation
The DL model was trained on a synthetically generated dataset of 3000 ECG samples with controlled variations in key morphological and temporal features within physiologically plausible ranges. A rule-based validation process discarded samples violating electrophysiological constraints to ensure internal consistency. Although the exclusive use of synthetic data limits clinical applicability, it supports the system’s educational goals by enabling reproducible, privacy-preserving training and alleviating common issues such as class imbalance and limited data availability.
The correctness of the generated ECG data was verified through a multi-level validation process focused on physiological plausibility and educational suitability. This process combined rule-based constraints to enforce clinically accepted ranges for heart rate, waveform durations, and temporal relationships, with morphological coherence checks to ensure the presence and continuity of the P wave, QRS complex, and T wave. In addition, representative generated signals were reviewed by cardiology experts to confirm their realism and usefulness for ECG education.
3.2 Model Selection
To select the most suitable ECG classification model, we evaluated logistic regression, support vector machines, random forest, and a multilayer perceptron using repeated stratified 5-fold cross-validation. Table 2 reports average accuracy, F1-score, and ROC-AUC across folds. While all models achieved competitive performance, the MLP obtained slightly lower scores than the traditional classifiers.
Table 2 Cross-validated performance (mean std) across repeated stratified k-folds
| Model | Accuracy | F1 | ROC-AUC |
| Logistic regression | 0.843 0.016 | 0.843 0.016 | 0.973 0.004 |
| Neural network (MLP) | 0.820 0.024 | 0.819 0.025 | 0.965 0.005 |
| Random forest | 0.839 0.019 | 0.838 0.020 | 0.970 0.004 |
| SVM (RBF) | 0.840 0.020 | 0.840 0.019 | 0.970 0.004 |
To determine whether these differences were statistically meaningful, we computed effect sizes between the MLP and each baseline using Cliff’s delta, mean difference, confidence intervals, and the statistic, a non-parametric measure of stochastic dominance between two distributions (where 0.5 indicates no difference), as shown in Table 3.
Table 3 Effect sizes comparing MLP and baselines (mean diff. MLPbaseline)
| Baseline | Metric | MLP mean | Diff | 95% CI | |
| Logistic regression | F1 | 0.819 | 0.023 | [0.031, 0.015] | 0.202 |
| Logistic regression | Accuracy | 0.820 | 0.023 | [0.031, 0.015] | 0.205 |
| SVM (RBF) | F1 | 0.819 | 0.021 | [0.027, 0.015] | 0.231 |
| SVM (RBF) | Accuracy | 0.820 | 0.020 | [0.027, 0.014] | 0.231 |
| Random forest | F1 | 0.819 | 0.019 | [0.026, 0.012] | 0.246 |
| Random forest | Accuracy | 0.820 | 0.019 | [0.027, 0.012] | 0.232 |
Although not the top performer, the MLP was selected for ECGTwinMentor due to real-time, edge-friendly inference and seamless simulator integration. Its modular design enables easy retraining with additional real or synthetic ECG data, aligning with the platform’s educational and adaptive goals. Given these practical benefits, the modest performance trade-off was deemed acceptable.
3.3 Model Performance
The model showed stable learning dynamics, with loss converging below 0.6 and validation accuracy stabilizing around 70–75%. On the test set, it achieved 86% accuracy, an MAE of 0.08, and an of 0.85, with a specificity of 94% for arrhythmias. These results confirm its predictive reliability and educational value. Inference times were below 50 ms on a desktop, 150 ms on an Android, and 500 ms on Raspberry Pi, enabling real-time interaction [10].
4 Experimental Setup
This work investigates the viability of integrating an interactive ECG educational platform with digital twin capabilities, deployable across various devices and settings. To evaluate this, ECGTwinMentor was implemented and tested in four representative environments:
• Web-based application: Developed using Python (Flask), TensorFlow, and HTML/CSS/JS for interactive visualization. Runs on standard desktop/laptop browsers (Chrome/Firefox).
• Raspberry Pi: Deployed on Raspberry Pi 4B with 4 GB RAM, using a minimal Flask-based backend and pre-loaded TFLite models for offline prediction.
• Android application: Built with Android Studio using Kotlin. Models were converted to TensorFlow Lite format for local on-device inference.
All experiments were conducted using synthetic ECG data generated with physiologically constrained parameters. The same trained neural network model was used across all platforms to ensure consistency in predictions. Device-specific optimizations (e.g., model quantization or hardware acceleration) were applied when needed to meet real-time interaction requirements.
4.1 ECGTwinMentor in the Cloud: Web-based Application
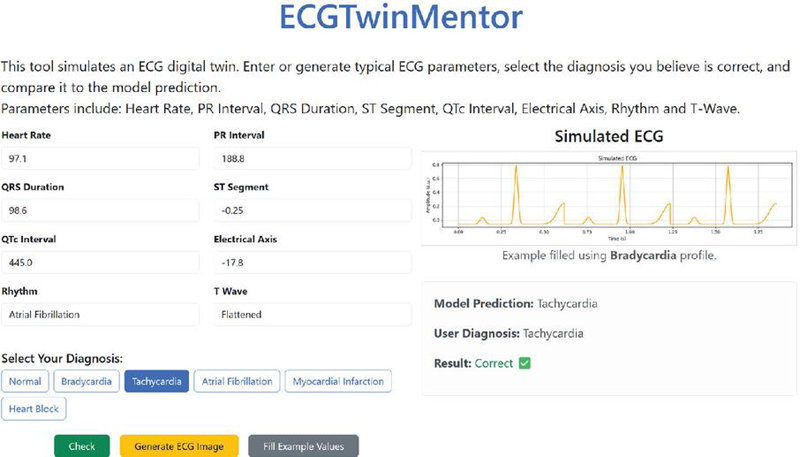
ECGTwinMentor includes a web application that supports ECG interpretation and diagnosis by allowing students to generate parameterized ECG simulations and verify their diagnoses using the DL model. The interactive simulator enables real-time parameter adjustment and waveform visualization in a risk-free setting, fostering diagnostic reasoning through hands-on practice. Integrated within a DL-based web platform, this module provides flexible and accessible training, as illustrated in Figure 3.
Figure 3 ECGTwinMentor web user interface.
4.1.1 Security enhancements
ECGTwinMentor incorporates several security mechanisms across both frontend and backend components. A restrictive cross-origin resource sharing (CORS) policy limits unauthorized cross-domain requests, while API rate limiting (100 requests per minute per IP) helps protect against abuse and denial-of-service (DoS) attacks. In addition, ECG inputs are encrypted on the client side using AES in CBC mode and securely transmitted to the server. Together, these measures enhance confidentiality, integrity, and resilience, ensuring a robust and privacy-aware platform for educational use.
4.2 ECGTwinMentor in the Edge: Low-consumption Devices
ECGTwinMentor is designed not only as a web-based educational platform but also for integration with edge computing environments. The system can operate using a local model, with optional updates from a remote server, enabling secure and offline use. To illustrate this capability, two edge-device implementations were developed, demonstrating how ECGTwinMentor can be extended beyond web interfaces to embedded and mobile systems.
4.2.1 Raspberry Pi deployment
The first edge implementation targets a Raspberry Pi, providing a minimal and cost-effective setup for ECG inference. A Python-based script performs on-device predictions using a TensorFlow Lite model via the tflite_runtime interpreter, with ECG parameters supplied through command-line arguments. The classification logic mirrors the web-based model and includes an update command to retrieve the latest model from a remote server, as illustrated in Listing 1.
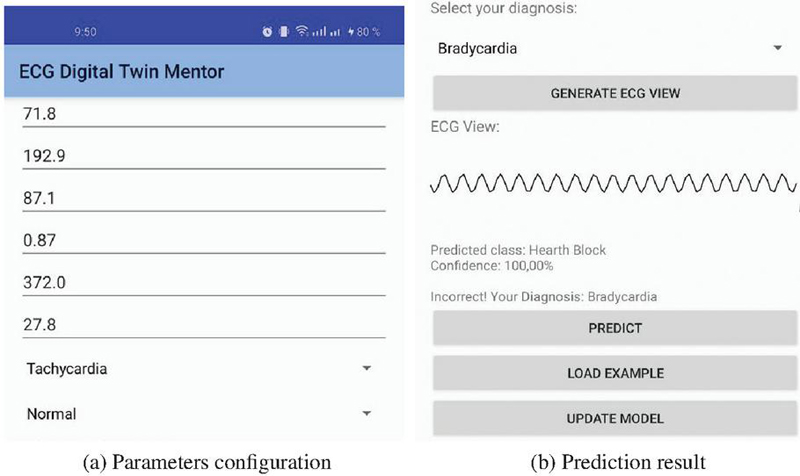
Figure 4 ECGTwinMentor Android implementation.
The Raspberry Pi used for testing was a Raspberry Pi 4 Model B, equipped with a quad-core ARM Cortex-A72 processor and 4 GB of LPDDR4 RAM. The device operated under a 64-bit Debian-based Raspberry Pi OS and utilized CPU-only inference with TensorFlow Lite. Despite its limited hardware, the Raspberry Pi achieved acceptable performance for real-time ECG predictions, with an average inference time below 500 milliseconds.
4.2.2 Android deployment
The second edge deployment is an Android application. This mobile version of ECGTwinMentor brings the system to smartphones and tablets, making it especially useful in educational settings or areas with limited infrastructure. The application integrates an embedded TensorFlow Lite model, enabling fast on-device predictions without the need for continuous server communication. The platform includes a self-assessment mechanism whereby users can select a suspected diagnosis and compare it against the model’s prediction. Also, the system supports model updates of the TFLite model from a remote server, ensuring adaptability and continuous improvement in training accuracy.
Input: User parameters ; physiological constraints ; ECG generator ; preprocessing ; trained classifier ; feedback rules ; max steps .
Output: Session log (parameters, ECG, validation, prediction, feedback).
1 ;
2 for to do
3 Receive updated parameters from user;
4 ;
5 // range + dependency checks
6 ;
7 // generate synthetic ECG
8 ;
// morphology/timing/artifact checks
9 if is invalid then
10 Display warning and request parameter revision;
11 Append to ;
continue;
12 ;
// normalize/resample/encode for model input
13 ;
// prediction + confidence
14 ;
15 // explanations + highlights
26 RENDER;
17 Append to ;
if user ends session then
break;
18 return ;
Algorithm 1 Pseudocode of ECGTwinMentor: reproducible flow.
For mobile deployment, the system was tested on a mid-range Android device, specifically a OnePlus 9, powered by a Qualcomm Snapdragon 888, 8 GB of RAM, an Adreno 660 GPU, and Android 14. The mobile version provided near-instantaneous inference, typically under 150 milliseconds, enabling a smooth user experience without relying on cloud resources.
These scenarios highlight the flexibility of ECGTwinMentor and its potential to adapt to various platforms and scenarios, maintaining consistency in predictive capability and educational value. ECGTwinMentor is available in the following repository for the deep learning model, the web, and edge devices 1.
4.3 Reproducibility
To improve clarity and reproducibility, Algorithm 1 summarizes the core workflow of the proposed ECGTwinMentor. The algorithm formalizes the experiential learning loop, starting from user-controlled parameter adjustment, followed by constrained ECG signal generation and physiological plausibility validation. The validated signal is then processed and fed into the trained deep learning model to obtain a diagnostic prediction and confidence estimates. Finally, pedagogical feedback and visual explanations are generated and presented to the learner, enabling iterative exploration of ECG patterns.
5 Validation
The validation aimed to assess both the technical and pedagogical effectiveness of ECGTwinMentor. Two complementary studies were conducted:
1. A survey with healthcare experts to evaluate realism, diagnostic accuracy, and clinical applicability.
2. A survey with medical students to assess usability, learning value, and engagement.
5.1 Validation with Healthcare Experts
On the one hand, six healthcare experts evaluated the realism, usability, and clinical coherence of ECGTwinMentor. The survey included nine quantitative items and one open-ended question. Table 4 summarizes the results.
Table 4 Summary of expert validation results (Likert 1–5; )
| Criterion | Mean | SD | % 4 |
| ECG simulation reflects real clinical patterns | 3.5 | 0.6 | 50% |
| Supports effective formative evaluation | 3.7 | 0.5 | 67% |
| Visual and dynamic representation is adequate | 4.5 | 0.8 | 83% |
| Design aligns with clinical workflow | 3.8 | 0.4 | 83% |
| Potential integration into medical programs | 4.0 | 0.6 | 83% |
| Parameter customization for different scenarios | 4.3 | 0.5 | 100% |
| Results consistent with real cases | 3.7 | 0.5 | 67% |
| Useful for research or specialized training | 4.2 | 0.4 | 100% |
| Overall system usability is satisfactory | 4.3 | 0.5 | 100% |
| Recommendation for professional use | 4.3 | 0.5 | 100% |
Experts rated ECGTwinMentor positively, particularly in terms of parameter customization, usability, and integration potential, with mean scores ranging from 3.5 to 4.5. Slightly lower ratings for realism and formative evaluation highlight areas for refinement, such as waveform variability and feedback mechanisms. Experts considered ECGTwinMentor realistic, pedagogically coherent, and suitable for educational and professional training.
5.2 Validation with Medical Students
On the other hand, eight medical students (4–5th year) evaluated usability, learning impact, motivation, and satisfaction. Table 5 presents the results.
Table 5 Summary of student validation results (Likert 1–5; )
| Criterion | Mean | SD | % 4 |
| Ease of use and understanding | 4.6 | 0.5 | 100% |
| Interface design is intuitive | 4.8 | 0.5 | 100% |
| Improved understanding of ECGs | 4.1 | 0.8 | 75% |
| Increased confidence interpreting ECGs | 4.4 | 0.7 | 88% |
| Simulation experience was motivating | 5.0 | 0.0 | 100% |
| Simulation reflects real learning scenarios | 5.0 | 0.0 | 100% |
| Facilitated active learning | 4.4 | 0.7 | 88% |
| Useful for peers at same academic level | 5.0 | 0.0 | 100% |
| Overall satisfaction with experience | 4.5 | 0.5 | 100% |
| Recommendation for academic training | 4.9 | 0.4 | 100% |
Students rated the tool highly across all categories, with the highest scores in motivation (5.0) and realism (5.0). Usability (4.6) and intuitiveness (4.8) confirm that the interface is accessible and well-designed. Satisfaction and recommendation rates reached nearly unanimous approval (4 for 100% of participants).
5.3 Comparative Analysis and Discussion
Both experts and students provided positive evaluations, confirming that ECGTwinMentor balances clinical realism and educational usability. Experts emphasized technical adequacy and customization potential, while students highlighted motivation, intuitive interaction, and learning impact. Overall, these results support ECGTwinMentor as an effective educational simulator suitable for integration into blended medical and cardiology training.
5.4 Threats to Validity
As ECGTwinMentor is an immersive educational digital twin, several validity threats must be considered. Internal validity may be affected by the use of simulated data generated under controlled conditions, while construct validity is limited by the abstraction level of the simulator, which may not capture all real-world ECG variability. External validity is constrained by the educational context and participant sample, and conclusion validity may be influenced by the selected evaluation metrics and exposure duration. Importantly, the system is intended for educational support and pattern exploration, not for clinical decision-making.
6 Related Work
Recent work on ECG interpretation and training leverages AI and new pedagogical paradigms. Notably, ECG-Image-Kit [23] generates multi-lead ECG images to support digitization and classification pipelines. While valuable for dataset construction, it lacks an interactive interface and integration in teaching environments, limiting its applicability to medical education.
Another relevant work is the digitization of paper-based ECGs, as proposed by Wu et al. [28]. Their deep learning-based approach allows the automated transformation of scanned ECG sheets into digital signals, thus facilitating data reuse and analysis. While highly relevant for archival and clinical purposes, this contribution does not address the pedagogical needs of students, nor does it provide feedback that could enhance training.
From the perspective of educational methodology, Wen et al. [27] apply the BOPPPS-CBL model (bridge-in, objective, pre-assessment, participatory learning, post-assessment, and summary) in ECG teaching for nursing students. This work demonstrates that a structured, learner-centered approach can significantly improve the acquisition of ECG interpretation. Nonetheless, it does not exploit the potential of artificial intelligence or immersive technologies, and remains limited to traditional classroom-based instruction.
In contrast, Qi and Su [18] propose a Cybertwin-based multimodal network for ECG monitoring in real time. Their system integrates deep learning with human activity recognition to improve continuous pattern monitoring, emphasizing scalability and robustness in clinical contexts. While this work showcases strong technical innovation, its focus lies in monitoring and diagnostics rather than in training or education.
Against this backdrop, ECGTwinMentor integrates physiologically constrained ECG simulation, deep learning-based diagnosis, and multi-platform deployment into a single system for cardiology education. Its lightweight design enables configurable, feedback-rich practice in a risk-free setting, including offline use on edge and mobile devices.
Table 6 provides a comparison highlighting the main purposes, methods, limitations, and differences with respect to ECGTwinMentor. Prior studies typically emphasize one dimension, such as data generation, digitization, or pedagogy, whereas ECGTwinMentor combines all into a unified digital twin.
Table 6 Comparison of ECGTwinMentor with current works
| Work/ref. | Purpose | Method | Data | Comment |
| ECGTwinMentor (this work) | ECG training with DT + DL | Simulation + DL (TFLite) | Synth. | Interactive sim., auto-diagnosis, edge/cloud deployment |
| ECG-Image-Kit [23] | Generate ECG imgs w/ artifacts | Time-series to image | Synth. | No UI; ECGTM adds interactivity and learning focus |
| Wu et al. [28] | Digitize paper ECGs | DL digitization | Real (scans) | ECGTM focuses on training, not digitization |
| Wen et al. [27] | Pedagogical ECG training | Structured teaching | Real | ECGTM adds AI, feedback, simulation and mobility |
| Qi and Su [18] | Real-time ECG monitoring | Cybertwin + CNN | Real (cont.) | ECGTM focuses on education, not monitoring |
7 Discussion
This section addresses the RQs based on the system’s design, validation, and preliminary evaluations.
• RQ1: Can simulated ECGs achieve sufficient realism for educational use? Yes. The simulator accurately replicates key waveform characteristics, and comparisons with clinical data support its realism for training purposes.
• RQ2: Does interactive use of the twin enhance recognition of patterns and abnormalities? Yes. Real-time manipulation of ECG parameters helps students understand how changes affect morphology, improving recognition of arrhythmias in tests.
• RQ3: How does predictive feedback influence learning outcomes and engagement? Immediate DL-based diagnostic suggestions allow students to reflect on their interpretations, enhancing both accuracy and motivation.
Deploying digital twins across cloud and edge environments involves trade-offs between resource constraints and interactivity. While edge execution offers low latency and improved responsiveness, it is limited by computational and energy resources; cloud deployment provides scalability for training and storage but depends on network connectivity.
Some limitations should be noted. The use of synthetic data may limit external validity, and the evaluation did not assess long-term learning outcomes or broader user contexts. Nevertheless, ECGTwinMentor provides a solid foundation for integrating AI-driven simulation into medical education, with future work focusing on expanded studies and pedagogical assessment.
8 Conclusions
ECGTwinMentor addresses key challenges in cardiology education by combining configurable ECG simulation, interactive practice, and immediate feedback in a safe training environment. The platform’s deep-learning model shows strong performance for automated cardiopathy prediction, while the simulator provides hands-on experience that strengthens practical ECG interpretation skills. Future work will extend validation to larger cohorts and incorporate longitudinal performance tracking to quantify learning gains, alongside improvements to model accuracy, an expanded ECG corpus, and deeper immersive features.
Acknowledgement
This work was supported by PID2021-124054OB-C31, TED2021-130913B-I00, and PDC2022-133465-I00 (MCIU/AEI/FEDER, UE); the Department of Economy, Science and Digital Agenda of the Government of Extremadura (GR24099, IB20094); the European Regional Development Fund; INCIBE; the “European Union NextGenerationEU/PRTR” (C110.23); and 0289_SER65_PLUS_6_P, co-financed by the European Union through the Interreg VI-A Spain-Portugal Program 2021-2027. We thank the COMPUTAEX Foundation and the Extremadura Supercomputing Center for access to the LUSITANIA supercomputer.
Footnotes
1This work is an extension of a previous paper: Flores-Martin, D.; Díaz-Barrancas, F.; Pardo, Pedro J.; Berrocal, J.; Murillo, J.M. ECGTwinMentor: Enhancing Cardiology Education in ECG with Digital Twins. The 5th International Workshop on Big data driven Edge Cloud Services (BECS 2025) Co-located with the 25th International Conference on Web Engineering (ICWE 2025), June 30-July 3, 2025, Delft, Netherlands.
1https://github.com/dflores0806/ECGVRDT
References
[1] Alazab, M., Khan, L.U., Koppu, S., Ramu, S.P., Boobalan, P., Baker, T., Maddikunta, P.K.R., Gadekallu, T.R., Aljuhani, A., et al.: Digital twins for healthcare 4.0-recent advances, architecture, and open challenges. IEEE Consumer Electronics Magazine 12(6), 29–37 (2022)
[2] Ali, O.M.A., Kareem, S.W., Mohammed, A.S.: Evaluation of electrocardiogram signals classification using cnn, svm, and lstm algorithm: A review. In: 2022 8th International Engineering Conference on Sustainable Technology and Development (IEC). pp. 185–191. IEEE (2022)
[3] Attia, Z.I., Harmon, D.M., Behr, E.R., Friedman, P.A.: Application of artificial intelligence to the electrocardiogram. European heart journal 42(46), 4717–4730 (2021)
[4] Avula, V., Wu, K.C., Carrick, R.T.: Clinical applications, methodology, and scientific reporting of electrocardiogram deep-learning models: a systematic review. JACC: Advances 2(10), 100686 (2023)
[5] Breen, C., Kelly, G., Kernohan, W.: Ecg interpretation skill acquisition: A review of learning, teaching and assessment. Journal of electrocardiology 73, 125–128 (2022)
[6] Chiu, T.K., Xia, Q., Zhou, X., Chai, C.S., Cheng, M.: Systematic literature review on opportunities, challenges, and future research recommendations of artificial intelligence in education. Computers and Education: Artificial Intelligence 4, 100118 (2023)
[7] Cluitmans, M.J., Plank, G., Heijman, J.: Digital twins for cardiac electrophysiology: state of the art and future challenges. Herzschrittmachertherapie+ Elektrophysiologie 35(2), 118–123 (2024)
[8] Cooper, J.B., Taqueti, V.: A brief history of the development of mannequin simulators for clinical education and training. Postgraduate medical journal 84(997), 563–570 (2008)
[9] Díaz-Barrancas, F., Flores-Martin, D., Berrocal, J., Peguero, J.C., Pardo, P.J.: Integrating real-time ecg data into virtual reality for enhanced medical training. In: 2025 IEEE Conference on Virtual Reality and 3D User Interfaces Abstracts and Workshops (VRW). pp. 1436–1437. IEEE (2025)
[10] Flores-Martin, D., Berrocal, J., García-Alonso, J., Murillo, J.M.: Extending w3c thing description to provide support for interactions of things in real-time. In: International conference on web engineering. pp. 30–41. Springer (2020)
[11] Flores-Martin, D., Laso, S., Berrocal, J., Murillo, J.M.: Towards digital health: Integrating federated learning and crowdsensing through the contigo app. SoftwareX 28, 101885 (2024)
[12] Grandits, T., Gillette, K., Plank, G., Pezzuto, S.: Accurate and efficient cardiac digital twin from surface ecgs: Insights into identifiability of ventricular conduction system. Medical Image Analysis 105, 103641 (2025). , https://www.sciencedirect.com/science/article/pii/S1361841525001884
[13] Iqbal, N., Kandasamy, R., Jyothish, K., Johnson, O., Sundaram, B., et al.: Virtual reality simulation for the acquisition and retention of electrocardiogram interpretation skills: A randomized controlled trial among undergraduate medical students. Cureus 16(6) (2024)
[14] Ko, Y., Issenberg, S.B., Roh, Y.S.: Effects of peer learning on nursing students’ learning outcomes in electrocardiogram education. Nurse education today 108, 105182 (2022)
[15] Li, C., Wu, Y., Lin, H., Li, J., Zhang, F., Yang, Y.: Ecg denoising method based on an improved vmd algorithm. IEEE Sensors Journal 22(23), 22725–22733 (2022)
[16] Park, J., An, J., Kim, J., Jung, S., Gil, Y., Jang, Y., Lee, K., Oh, I.y.: Study on the use of standard 12-lead ecg data for rhythm-type ecg classification problems. Computer Methods and Programs in Biomedicine 214, 106521 (2022)
[17] Patro, K.K., Jaya Prakash, A., Jayamanmadha Rao, M., Rajesh Kumar, P.: An efficient optimized feature selection with machine learning approach for ecg biometric recognition. IETE Journal of Research 68(4), 2743–2754 (2022)
[18] Qi, W., Su, H.: A cybertwin based multimodal network for ecg patterns monitoring using deep learning. IEEE Transactions on Industrial Informatics 18(10), 6663–6670 (2022)
[19] Qian, S., Ugurlu, D., Fairweather, E., Toso, L.D., Deng, Y., Strocchi, M., Cicci, L., Jones, R.E., Zaidi, H., Prasad, S., et al.: Developing cardiac digital twin populations powered by machine learning provides electrophysiological insights in conduction and repolarization. Nature Cardiovascular Research 4(5), 624–636 (2025)
[20] Rafie, N., Kashou, A.H., Noseworthy, P.A.: Ecg interpretation: clinical relevance, challenges, and advances. Hearts 2(4), 505–513 (2021)
[21] Rijnbeek, P.R., Van Herpen, G., Bots, M.L., Man, S., Verweij, N., Hofman, A., Hillege, H., Numans, M.E., Swenne, C.A., Witteman, J.C., et al.: Normal values of the electrocardiogram for ages 16–90 years. Journal of electrocardiology 47(6) (2014)
[22] Salvador, M., Kong, F., Peirlinck, M., Parker, D.W., Chubb, H., Dubin, A.M., Marsden, A.L.: Digital twinning of cardiac electrophysiology for congenital heart disease. Journal of the Royal Society Interface 21(215), 20230729 (2024)
[23] Shivashankara, K.K., Shervedani, A.M., Clifford, G.D., Reyna, M.A., Sameni, R., et al.: Ecg-image-kit: a synthetic image generation toolbox to facilitate deep learning-based electrocardiogram digitization. Physiological measurement 45(5), 055019 (2024)
[24] Siontis, K.C., Noseworthy, P.A., Attia, Z.I., Friedman, P.A.: Artificial intelligence-enhanced electrocardiography in cardiovascular disease management. Nature Reviews Cardiology 18(7), 465–478 (2021)
[25] Somani, S., Russak, A.J., Richter, F., Zhao, S., Vaid, A., Chaudhry, F., De Freitas, J.K., Naik, N., Miotto, R., Nadkarni, G.N., et al.: Deep learning and the electrocardiogram: review of the current state-of-the-art. EP Europace 23(8), 1179–1191 (2021)
[26] Stamate, E., Piraianu, A.I., Ciobotaru, O.R., Crassas, R., Duca, O., Fulga, A., Grigore, I., Vintila, V., Fulga, I., Ciobotaru, O.C.: Revolutionizing cardiology through artificial intelligence—big data from proactive prevention to precise diagnostics and cutting-edge treatment—a comprehensive review of the past 5 years. Diagnostics 14(11), 1103 (2024)
[27] Wen, H., Xu, W., Chen, F., Jiang, X., Zhang, R., Zeng, J., Peng, L., Chen, Y.: Application of the boppps-cbl model in electrocardiogram teaching for nursing students: a randomized comparison. BMC Medical Education 23(1), 987 (2023)
[28] Wu, H., Patel, K.H.K., Li, X., Zhang, B., Galazis, C., Bajaj, N., Sau, A., Shi, X., Sun, L., Tao, Y., et al.: A fully-automated paper ecg digitisation algorithm using deep learning. Scientific Reports 12(1), 20963 (2022)
Biographies

Daniel Flores-Martin is a researcher and head of the systems and supercomputing department at COMPUTAEX. His research interests include the Internet of Things, artificial intelligence, and high-performance computing.

Francisco Díaz-Barrancas is a post-doc researcher at the University of Extremadura. His main interests are virtual reality, artificial intelligence and color processing.

Pedro J. Pardo is an Associate Professor at the University of Extremadura. His research interests include color vision, neural networks and computer networks.

Javier Berrocal (IEEE Member) is an Associate Professor at the University of Extremadura. His main research interests are software architectures, mobile computing, and edge and fog computing.

Juan Manuel Murillo (IEEE Member) is a Full Professor at the University of Extremadura. His research interests include software architectures, mobile computing, and cloud computing.
Journal of Web Engineering, Vol. 25_3, 373–394
doi: 10.13052/jwe1540-9589.2534
© 2026 River Publishers