Bioinformatics Analyses of Oct4 Pseudogenes
DOI:
https://doi.org/10.13052/ijts2246-8765.2025.024Keywords:
Stem cell, pseudogenes, Octamer 4, microRNAAbstract
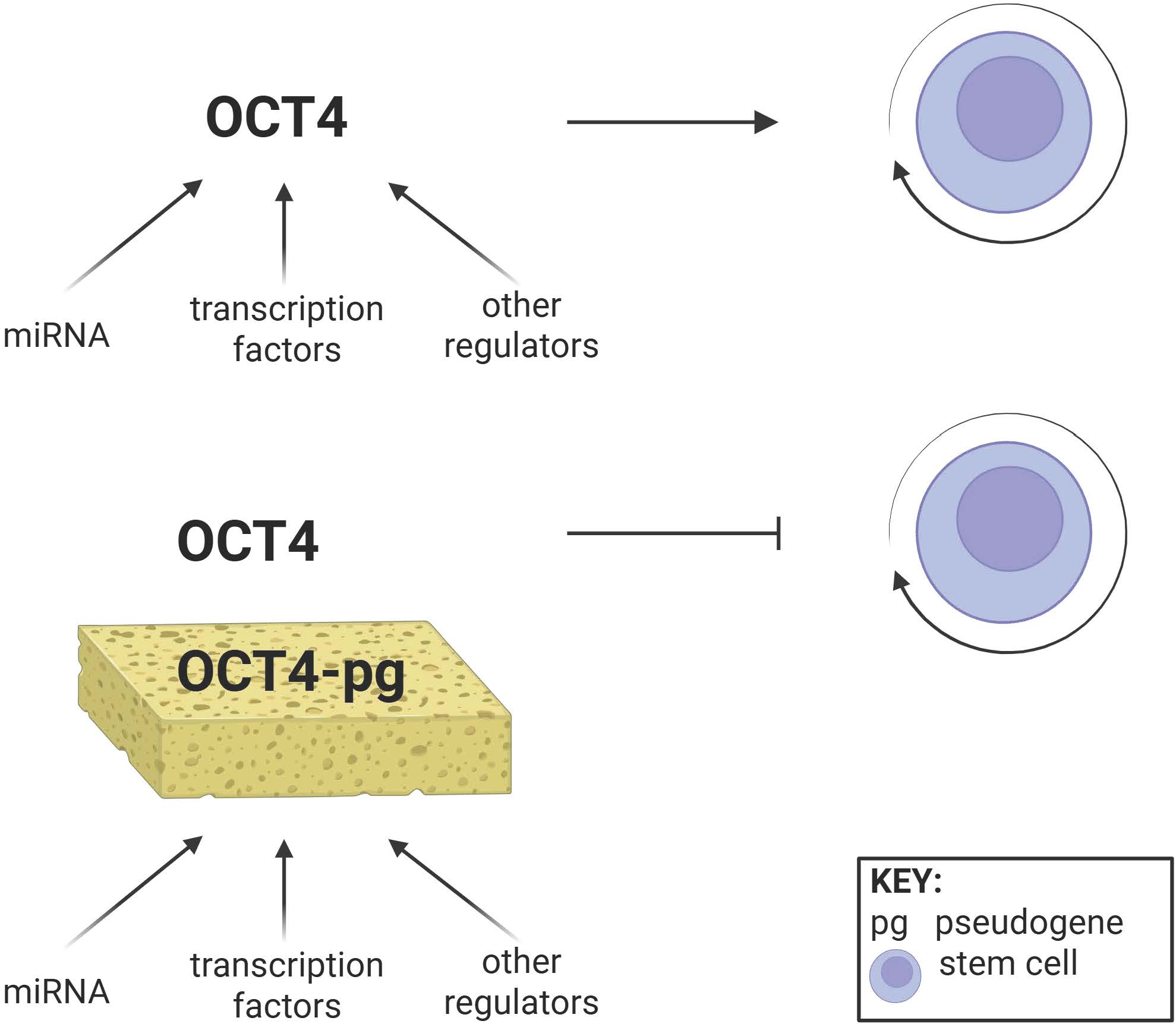
Oct4 is among the key genes involved in pluripotency. Oct4, through alternative splicing, produces at least three different transcripts, Oct4a, Oct4b, and Oct4b1. The Oct4a transcript has been focused on due to its role in pluripotency and multipotency of stem cells. There are several Oct4 pseudogenes with at least seven identified in the human genome. Some of these Oct4 pseudogenes have been found in various cancers. The Oct4 pseudogenes share high homology to the Oct4a transcript. There is limited information on the biologic role of Oct4 pseudogenes. More importantly, it is unclear how these pseudogenes affect the biology of the full-length transcript. Thus, it is important to understand these pseudogenes since their biology could lead to a better understanding of differentiation, cancer, and dedifferentiation to form cancer stem cells. This brief report used in silico analyses to analyze Oct4 and its pseudogenes. The implication for the findings on stem cells and cancer, as well as other related genes, are discussed.
Downloads
References
Wang, L., Z.-Y. Guo, R. Zhang, B. Xin, R. Chen, J. Zhao, T. Wang, W.-H. Wen, L.-T. Jia, and L.-B. Yao. 2013. Pseudogene OCT4-pg4 functions as a natural micro RNA sponge to regulate OCT4 expression by competing for miR-145 in hepatocellular carcinoma. Carcinogenesis 34: 1773–1781.
Suo, G., J. Han, X. Wang, J. Zhang, Y. Zhao, Y. Zhao, and J. Dai. 2005. Oct4 pseudogenes are transcribed in cancers. Biochem Biophysical Res Commun 337: 1047–1051.
Kolenda, T., P. Chałaj, A. Cichowicz, A. Trojańska, A. Bałoniak, M. Kwaśniewska, M. Odrobińska, K. Guglas, J. Kozłowska-Masłoń, P. Gieremek, P. Poter, M. Janiczek-Polewska, A. Florczak-Substyk, A. Przybyła, P. Mantaj, K. Regulska, B. J. Stanisz, Z. Cybulski, and U. Kazimierczak. 2025. Pseudogenes in the carcinogenesis: epithelial-to-mesenchymal transition process and cancer initiating cells. Rep Pract Oncol Radiother 30: 366–384.
Sharma, D., S. Gupta, G. Koshy, V. K. Sharma, M. Kamboj, and A. Hooda. 2025. Expression profile of cancer stem cell markers SOX2, OCT4 & NANOG in salivary gland malignancies: A systematic review. Indian J Med Res 161: 636–646.
Bliss, S. A., G. Sinha, O. A. Sandiford, L. M. Williams, D. J. Engelberth, K. Guiro, L. L. Isenalumhe, S. J. Greco, S. Ayer, M. Bryan, R. Kumar, N. M. Ponzio, and P. Rameshwar. 2016. Mesenchymal Stem Cell-Derived Exosomes Stimulate Cycling Quiescence and Early Breast Cancer Dormancy in Bone Marrow. Cancer Res 76: 5832–5844.
Sinha, G., A. I. Ferrer, S. Ayer, M. H. El-Far, S. H. Pamarthi, Y. Naaldijk, P. Barak, O. A. Sandiford, B. M. Bibber, G. Yehia, S. J. Greco, J. G. Jiang, M. Bryan, R. Kumar, N. M. Ponzio, J. P. Etchegaray, and P. Rameshwar. 2021. Specific N-cadherin-dependent pathways drive human breast cancer dormancy in bone marrow. Life Sci Alliance 4.
McGinnis, S., and T. L. Madden. 2004. BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res 32: W20–W25.
Li, W., A. Cowley, M. Uludag, T. Gur, H. McWilliam, S. Squizzato, Y. M. Park, N. Buso, and R. Lopez. 2015. The EMBL-EBI bioinformatics web and programmatic tools framework. Nucleic Acids Res 43: W580–W584.
McWilliam, H., W. Li, M. Uludag, S. Squizzato, Y. M. Park, N. Buso, A. P. Cowley, and R. Lopez. 2013. Analysis tool web services from the EMBL-EBI. Nucleic Acids Res 41: W597–W600.
Sievers, F., A. Wilm, D. Dineen, T. J. Gibson, K. Karplus, W. Li, R. Lopez, H. McWilliam, M. Remmert, and J. Söding. 2011. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Systems Biol 7: 539.
Yates, A., W. Akanni, M. R. Amode, D. Barrell, K. Billis, D. Carvalho-Silva, C. Cummins, P. Clapham, S. Fitzgerald, L. Gil, C. G. Girón, L. Gordon, T. Hourlier, S. E. Hunt, S. H. Janacek, N. Johnson, T. Juettemann, S. Keenan, I. Lavidas, F. J. Martin, T. Maurel, W. McLaren, D. N. Murphy, R. Nag, M. Nuhn, A. Parker, M. Patricio, M. Pignatelli, M. Rahtz, H. S. Riat, D. Sheppard, K. Taylor, A. Thormann, A. Vullo, S. P. Wilder, A. Zadissa, E. Birney, J. Harrow, M. Muffato, E. Perry, M. Ruffier, G. Spudich, S. J. Trevanion, F. Cunningham, B. L. Aken, D. R. Zerbino, and P. Flicek. 2016. Ensembl 2016. Nucleic Acids Res 44: D710–716.
Wang, X., and J. Dai. 2010. Concise review: isoforms of OCT4 contribute to the confusing diversity in stem cell biology. Stem Cells 28: 885–893.
Patel, S. A., S. H. Ramkissoon, M. Bryan, L. F. Pliner, G. Dontu, P. S. Patel, S. Amiri, S. R. Pine, and P. Rameshwar. 2012. Delineation of breast cancer cell hierarchy identifies the subset responsible for dormancy. Sci Rep 2: 906.
Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J Mol Biol 215: 403–410.
Poursani, E. M., B. M. Soltani, and S. J. Mowla. 2016. Differential expression of OCT4 pseudogenes in pluripotent and tumor cell lines. Cell J (Yakhteh) 18: 28.
Greco, S. J., K. Liu, and P. Rameshwar. 2007. Functional similarities among genes regulated by OCT4 in human mesenchymal and embryonic stem cells. Stem Cells 25: 3143–3154.
Kulcheski, F. R., A. P. Christoff, and R. Margis. 2016. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J Biotechnol 238: 42–51.
Villodre, E. S., F. C. Kipper, M. B. Pereira, and G. Lenz. 2016. Roles of OCT4 in tumorigenesis, cancer therapy resistance and prognosis. Cancer treatment reviews 51: 1–9.
Zhang, Q., Z. Han, Y. Zhu, J. Chen, and W. Li. 2020. The role and specific mechanism of OCT4 in cancer stem cells: a review. Intl J Stem Cells 13: 312–325.
Booth, H. A. F., and P. W. Holland. 2004. Eleven daughters of NANOG. Genomics 84: 229–238.